This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Luke Edward Severinac/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 9: | Line 9: | ||
==Active State== | ==Active State== | ||
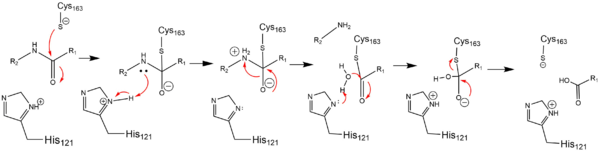
| - | In order to function as an endoprotease, Caspase-6 binds a <scene name='75/752344/Protein_ligand_real/1'>ligand</scene>, which can include neuronal proteins and [https://en.wikipedia.org/wiki/Tubulin tubulins], in its active site.[[Image:Binding grove active caspase 6.png|100 px|right|thumb|Substrate binding groove in Caspase-6. Blue - catalytic residues | + | In order to function as an endoprotease, Caspase-6 utilzes a catalytic triad that binds a <scene name='75/752344/Protein_ligand_real/1'>ligand</scene>, which can include neuronal proteins and [https://en.wikipedia.org/wiki/Tubulin tubulins], in its active site. This binding groove contains three critical amino acid residues necessary to perform cleavage of the peptide bonds. A <scene name='75/752344/Catalytic_triad_real/1'>catalytic triad</scene>, composed of <scene name='75/752344/His121_real/1'>His-121</scene>, <scene name='75/752344/Glu123_real/1'>Glu-123</scene>, and <scene name='75/752344/Cys163_real/1'>Cys-163</scene>, carries out cleavage of the substrate. In the theorized mechanism, His-121 acts as an acid catalyst, Glu-123 acts as a base catalyst to deprotonate Cys-163, which then acts as covalent catalyst. |
| + | [[Image:Cystine Aspartase.png|600 px|active site mechanism]] | ||
| + | [[Image:Binding grove active caspase 6.png|100 px|right|thumb|Substrate binding groove in Caspase-6. Blue - catalytic residues | ||
yellow - ligand | yellow - ligand | ||
| - | red - generic surface]] | + | red - generic surface]] |
| - | + | ||
==Zinc Inhibition== | ==Zinc Inhibition== | ||
Caspase-6 function is primarily inhibited by the binding of a <scene name='75/752344/Zinc_caspase-6/1'>zinc</scene> ion, which binds to an <scene name='75/752344/Caspase6_allosteric_site/1'>allosteric site</scene> instead of the <scene name='75/752344/Caspase6_allostericactiv_site/1'>active site</scene>. This allosteric site is located on the outside of the protein and is distal to the active site. The zinc ion is bound to <scene name='75/752344/Caspase6_allosteric_site_resid/1'>three amino acid residues</scene>, Lys-36, Glu-244, and His-287. Once the ion is bound to the protein, it is then stabilized by a <scene name='75/752344/H20_zinc_binding_casp/1'>water molecule</scene> found in the cytoplasm. The binding of zinc at the exosite is suggested to cause a conformational change in the protein from an <scene name='75/752344/Catalytic_triad_real/1'>active state</scene> to an <scene name='75/752344/Inactive_catalytic_triad_casp/1'>inactive state</scene> that misaligns catalytic residues and inhibits activity of the enzyme. It has been proposed that helices of the active dimer must rotate or move in some other way to provide these ideal interactions with zinc. This subtle shift is most likely the cause for allosteric inhibition. As the helices move to bind zinc, the amino acids of the active site become misaligned. The altered positions of the amino acids no longer provide ideal interactions for incoming substrates. After zinc binds, no new substrates enter the active site. Thus, Caspase-6 is effectively inhibited. The residues in the active site no longer provide ideal interactions with the substrate and therefore, substrate does not bind. Zinc binding to the exosite is tightly regulated as it inhibits Caspase-6's critical role in initiation of apoptosis. | Caspase-6 function is primarily inhibited by the binding of a <scene name='75/752344/Zinc_caspase-6/1'>zinc</scene> ion, which binds to an <scene name='75/752344/Caspase6_allosteric_site/1'>allosteric site</scene> instead of the <scene name='75/752344/Caspase6_allostericactiv_site/1'>active site</scene>. This allosteric site is located on the outside of the protein and is distal to the active site. The zinc ion is bound to <scene name='75/752344/Caspase6_allosteric_site_resid/1'>three amino acid residues</scene>, Lys-36, Glu-244, and His-287. Once the ion is bound to the protein, it is then stabilized by a <scene name='75/752344/H20_zinc_binding_casp/1'>water molecule</scene> found in the cytoplasm. The binding of zinc at the exosite is suggested to cause a conformational change in the protein from an <scene name='75/752344/Catalytic_triad_real/1'>active state</scene> to an <scene name='75/752344/Inactive_catalytic_triad_casp/1'>inactive state</scene> that misaligns catalytic residues and inhibits activity of the enzyme. It has been proposed that helices of the active dimer must rotate or move in some other way to provide these ideal interactions with zinc. This subtle shift is most likely the cause for allosteric inhibition. As the helices move to bind zinc, the amino acids of the active site become misaligned. The altered positions of the amino acids no longer provide ideal interactions for incoming substrates. After zinc binds, no new substrates enter the active site. Thus, Caspase-6 is effectively inhibited. The residues in the active site no longer provide ideal interactions with the substrate and therefore, substrate does not bind. Zinc binding to the exosite is tightly regulated as it inhibits Caspase-6's critical role in initiation of apoptosis. | ||
Revision as of 03:04, 20 April 2017
Caspase-6 in Homo sapiens
| |||||||||||