This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Luke Edward Severinac/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 8: | Line 8: | ||
In addition to a self-cleavage mechanism, Caspase-6 <scene name='75/752344/Caspase-6_zymogen_yeahboi/1'>zymogen</scene> can be activated getting cleaved by Caspase-3, as well as other enzymes. The mechanism of activation by clevage is highly conserved across the caspase family; Self-processing is uniquely recognized as the primary mechanism for Caspase-6 activation, where clevage must occur at <scene name='75/752344/Caspase-6_cleavage_sites_real/1'>three sites</scene> for complete activation, specifically the <scene name='75/752344/Caspase-6_prodomain/1'>pro-domain</scene> and the <scene name='75/752344/Caspase-6_intersubunit_linker/1'>intersubunit linker</scene> must be removed. These cleavages are both sequence specific and ordered, starting with cleavage of the pro-domain at <scene name='75/752344/Caspase-6_prodomain_cleavage/1'>residue 30</scene>. Then removal of the intersubunit linker occurs through cleavage at two sites, <scene name='75/752344/Caspase-6_176-179_cleavageyis/1'>DVVD179 and TEVD193</scene>. It has been proposed that this sequence of cleavage is due to the pro-domain being more readily available to enter the active site, whose presence inhibits Caspase-6's ability to cleave the intersubunit loop and self-activate; The prodomain acts as a “suicide protector”, preventing the TEVD193 cleavage site from the active site[3]. After both cleavages occur, <scene name='75/752344/Active_caspase_6_dimer/1'>active caspase-6</scene> remains in solution as a dimer. | In addition to a self-cleavage mechanism, Caspase-6 <scene name='75/752344/Caspase-6_zymogen_yeahboi/1'>zymogen</scene> can be activated getting cleaved by Caspase-3, as well as other enzymes. The mechanism of activation by clevage is highly conserved across the caspase family; Self-processing is uniquely recognized as the primary mechanism for Caspase-6 activation, where clevage must occur at <scene name='75/752344/Caspase-6_cleavage_sites_real/1'>three sites</scene> for complete activation, specifically the <scene name='75/752344/Caspase-6_prodomain/1'>pro-domain</scene> and the <scene name='75/752344/Caspase-6_intersubunit_linker/1'>intersubunit linker</scene> must be removed. These cleavages are both sequence specific and ordered, starting with cleavage of the pro-domain at <scene name='75/752344/Caspase-6_prodomain_cleavage/1'>residue 30</scene>. Then removal of the intersubunit linker occurs through cleavage at two sites, <scene name='75/752344/Caspase-6_176-179_cleavageyis/1'>DVVD179 and TEVD193</scene>. It has been proposed that this sequence of cleavage is due to the pro-domain being more readily available to enter the active site, whose presence inhibits Caspase-6's ability to cleave the intersubunit loop and self-activate; The prodomain acts as a “suicide protector”, preventing the TEVD193 cleavage site from the active site[3]. After both cleavages occur, <scene name='75/752344/Active_caspase_6_dimer/1'>active caspase-6</scene> remains in solution as a dimer. | ||
| - | ==Active State==[[Image:Binding grove active caspase 6.png|100 px|right|thumb|Substrate binding groove in Caspase-6. Blue - catalytic residues | + | ==Active State== |
| + | [[Image:Binding grove active caspase 6.png|100 px|right|thumb|Substrate binding groove in Caspase-6. Blue - catalytic residues | ||
yellow - ligand | yellow - ligand | ||
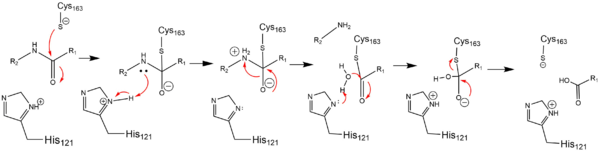
| - | red - generic surface]] | + | red - generic surface]]In order to function as an endoprotease, Caspase-6 utilzes a catalytic triad that binds a <scene name='75/752344/Protein_ligand_real/1'>ligand</scene>, which can include neuronal proteins and [https://en.wikipedia.org/wiki/Tubulin tubulins], in its active site. This binding groove contains three critical amino acid residues necessary to perform cleavage of the peptide bonds. A <scene name='75/752344/Catalytic_triad_real/1'>catalytic triad</scene>, composed of <scene name='75/752344/His121_real/1'>His-121</scene>, <scene name='75/752344/Glu123_real/1'>Glu-123</scene>, and <scene name='75/752344/Cys163_real/1'>Cys-163</scene>, carries out cleavage of the substrate. In the theorized mechanism, His-121 acts as an acid catalyst, Glu-123 acts as a base catalyst to deprotonate Cys-163, which then acts as covalent catalyst. |
| - | In order to function as an endoprotease, Caspase-6 utilzes a catalytic triad that binds a <scene name='75/752344/Protein_ligand_real/1'>ligand</scene>, which can include neuronal proteins and [https://en.wikipedia.org/wiki/Tubulin tubulins], in its active site. This binding groove contains three critical amino acid residues necessary to perform cleavage of the peptide bonds. A <scene name='75/752344/Catalytic_triad_real/1'>catalytic triad</scene>, composed of <scene name='75/752344/His121_real/1'>His-121</scene>, <scene name='75/752344/Glu123_real/1'>Glu-123</scene>, and <scene name='75/752344/Cys163_real/1'>Cys-163</scene>, carries out cleavage of the substrate. In the theorized mechanism, His-121 acts as an acid catalyst, Glu-123 acts as a base catalyst to deprotonate Cys-163, which then acts as covalent catalyst. | + | |
[[Image:Cystine Aspartase.png|600 px|active site mechanism]] | [[Image:Cystine Aspartase.png|600 px|active site mechanism]] | ||
Revision as of 03:15, 20 April 2017
Caspase-6 in Homo sapiens
| |||||||||||