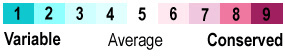User:Natalie Van Ochten/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 14: | Line 14: | ||
===Lid Region=== | ===Lid Region=== | ||
Amino acids 25-36 of DDAH constitute the flexible | Amino acids 25-36 of DDAH constitute the flexible | ||
| - | <scene name='75/752351/Lid_focus/1'>loop region</scene> of the protein which is more commonly known as the lid region <ref name="frey" />. The lid is what allows the active site to be exposed to substrate binding or not. Studies have shown crystal structures of the lid at <scene name='69/694225/Open_surface/5'>open</scene>and <scene name='69/694225/Closed_surface/4'>closed</scene>conformations. In the open conformation, the lid forms an alpha helix and the amino acid <scene name='69/694225/Lid_helix/1'>Leu29</scene> is moved so it does not interact with the active site. This allows the active site to be vulnerable to attack. When the lid is closed, a specific <scene name=’75/752351/Hbond_leu29/1’>hydrogen bond</scene> can form between the Leu29 carbonyl and the amino group on bound molecule. This stabilizes this complex. The Leu29 is then <scene name='69/694225/Closed_lid_zn9/3'>blocking</scene> the active site entrance <ref name="frey" />. Opening and closing the lid takes place faster than the actual reaction in the active site <ref name="rasheed">Rasheed M, Richter C, Chisty LT, Kirkpatrick J, Blackledge M, Webb MR, Driscoll PC. Ligand-dependent dynamics of the active site lid in bacterial Dimethyarginine Dimethylaminohydrolase. Biochemistry. 2014 Feb 18;53:1092-1104. PMCID:<span class="plainlinks">[https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3945819/ PMC3945819]</span> doi:<span class="plainlinks">[http://pubs.acs.org/doi/abs/10.1021/bi4015924 10.1021/bi4015924]</span></ref>. This suggests that the <span class="plainlinks">[https://en.wikipedia.org/wiki/Rate-determining_step rate-limiting step]</span> of this reaction is not the lid movement but is the actual chemistry happening to the substrate in the active site of DDAH <ref name="rasheed" />. | + | <scene name='75/752351/Lid_focus/1'>loop region</scene> of the protein which is more commonly known as the lid region <ref name="frey" />. The lid is what allows the active site to be exposed to substrate binding or not. Studies have shown crystal structures of the lid at <scene name='69/694225/Open_surface/5'>open</scene>and <scene name='69/694225/Closed_surface/4'>closed</scene> conformations. In the open conformation, the lid forms an alpha helix and the amino acid <scene name='69/694225/Lid_helix/1'>Leu29</scene> is moved so it does not interact with the active site. This allows the active site to be vulnerable to attack. When the lid is closed, a specific <scene name=’75/752351/Hbond_leu29/1’>hydrogen bond</scene> can form between the Leu29 carbonyl and the amino group on bound molecule. This stabilizes this complex. The Leu29 is then <scene name='69/694225/Closed_lid_zn9/3'>blocking</scene> the active site entrance <ref name="frey" />. Opening and closing the lid takes place faster than the actual reaction in the active site <ref name="rasheed">Rasheed M, Richter C, Chisty LT, Kirkpatrick J, Blackledge M, Webb MR, Driscoll PC. Ligand-dependent dynamics of the active site lid in bacterial Dimethyarginine Dimethylaminohydrolase. Biochemistry. 2014 Feb 18;53:1092-1104. PMCID:<span class="plainlinks">[https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3945819/ PMC3945819]</span> doi:<span class="plainlinks">[http://pubs.acs.org/doi/abs/10.1021/bi4015924 10.1021/bi4015924]</span></ref>. This suggests that the <span class="plainlinks">[https://en.wikipedia.org/wiki/Rate-determining_step rate-limiting step]</span> of this reaction is not the lid movement but is the actual chemistry happening to the substrate in the active site of DDAH <ref name="rasheed" />. |
====Lid Region Conservation==== | ====Lid Region Conservation==== | ||
Revision as of 00:56, 21 April 2017
Dimethylarginine Dimethylaminohydrolase
| |||||||||||
References
- ↑ 1.0 1.1 Palm F, Onozato ML, Luo Z, Wilcox CS. Dimethylarginine dimethylaminohydrolase (DDAH): expression, regulation, and function in the cardiovascular and renal systems. American Journal of Physiology. 2007 Dec 1;293(6):3227-3245. PMID:17933965 doi:10.1152/ajpheart.00998.2007
- ↑ 2.0 2.1 2.2 Tran CTL, Leiper JM, Vallance P. The DDAH/ADMA/NOS pathway. Atherosclerosis Supplements. 2003 Dec;4(4):33-40. PMID:14664901 doi:10.1016/S1567-5688(03)00032-1
- ↑ 3.00 3.01 3.02 3.03 3.04 3.05 3.06 3.07 3.08 3.09 3.10 3.11 3.12 3.13 3.14 3.15 3.16 3.17 3.18 3.19 3.20 3.21 3.22 3.23 Frey D, Braun O, Briand C, Vasak M, Grutter MG. Structure of the mammalian NOS regulator dimethylarginine dimethylaminohydrolase: a basis for the design of specific inhibitors. Structure. 2006 May;14(5):901-911. PMID:[1] doi:10.1016/j.str.2006.03.006
- ↑ Humm A, Fritsche E, Mann K, Göhl M, Huber R. Recombinant expression and isolation of human L-arginine:glycine amidinotransferase and identification of its active-site cysteine residue. Biochemical Journal. 1997 March 15;322(3):771-776. PMID:9148748 doi:10.1042/bj3220771
- ↑ 5.0 5.1 5.2 Rasheed M, Richter C, Chisty LT, Kirkpatrick J, Blackledge M, Webb MR, Driscoll PC. Ligand-dependent dynamics of the active site lid in bacterial Dimethyarginine Dimethylaminohydrolase. Biochemistry. 2014 Feb 18;53:1092-1104. PMCID:PMC3945819 doi:10.1021/bi4015924
- ↑ 6.0 6.1 Stone EM, Costello AL, Tierney DL, Fast W. Substrate-assisted cysteine deprotonation in the mechanism of Dimethylargininase (DDAH) from Pseudomonas aeruginosa. Biochemistry. 2006 May 2;45(17):5618-5630. PMID:16634643 doi:10.1021/bi052595m
- ↑ 7.0 7.1 Pace NJ, Weerpana E. Zinc-binding cysteines: diverse functions and structural motifs. Biomolecules. 2014 June;4(2):419-434. PMCID:4101490 doi:10.3390/biom4020419
- ↑ Janssen W, Pullamsetti SS, Cooke J, Weissmann N, Guenther A, Schermuly RT. The role of dimethylarginine dimethylaminohydrolase (DDAH) in pulmonary fibrosis. The Journal of Pathology. 2012 Dec 12;229(2):242-249. Epub 2013 Jan. PMID:23097221 doi:10.1002/path.4127
- ↑ Colasanti M, Suzuki H. The dual personality of NO. ScienceDirect. 2000 Jul 1;21(7):249-252. PMID:10979862 doi:10.1016/S0165-6147(00)01499-1
- ↑ Rassaf T, Feelisch M, Kelm M. Circulating NO pool: assessment of nitrite and nitroso species in blood and tissues. Free Rad. Biol. Med. 2004 Feb 15;36(4):413-422. PMID:14975444 doi:10.1016/j.freeradbiomed.2003.11.011
- ↑ Tsao PS, Cooke JP. Endothelial alterations in hypercholesterolemia: more than simply vasodilator dysfunction. Journal of Cardiovascular Pharmacology. 1998;32(3):48-53. PMID:9883748
- ↑ Vallance P, Leiper J. Blocking NO synthesis: how, where and why? Nat. Rev. Drug Discov. 2002 Dec;1(12):939-950. PMID:12461516 doi:10.1038/nrd960
Student Contributors
- Natalie Van Ochten
- Kaitlyn Enderle
- Colton Junod
3D Structures of Dimethylarginine Dimethylaminohydrolase
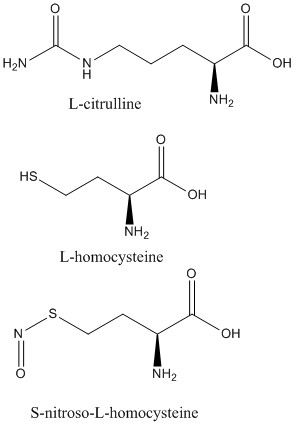
2C6Z L-citrulline bound to isoform 1
2CI1 S-nitroso-L-homocysteine bound to isoform 1
2CI3 crystal form 1
2CI4 crystal form 2
2CI5 L-homocysteine bound to isoform 1
2CI6 Zn (II) bound at low pH to isoform 1
2CI7 Zn (II) bound at high pH to isoform 1