This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Rhodopsin
From Proteopedia
(Difference between revisions)
| Line 23: | Line 23: | ||
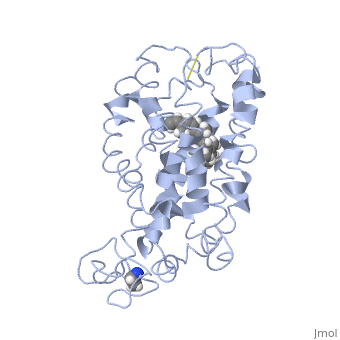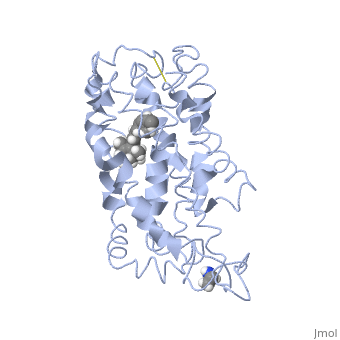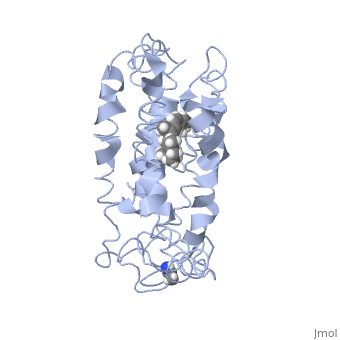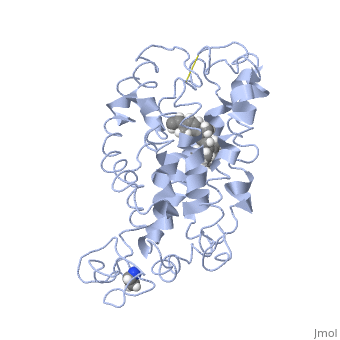
===Retinal Chromophore of Rhodopsin=== | ===Retinal Chromophore of Rhodopsin=== | ||
| - | Rhodopsin consists of an opsin [http://en.wikipedia.org/wiki/Apoprotein apoprotein] and a <scene name='40/400594/Cv/7'>11-cis retinylidene chromophore</scene> in its active site. Rhodopsin is bound covalently to the 11-''cis'' retinal, the chromophore or "ligand," (shown in <span style="color:yellow;background-color:black;font-weight:bold;">yellow</span>) and this retinal is found in deeply in the core of the helices, in a hydrophobic site, parallel to the lipid bilayer<ref name="Article19">PMID:16051215</ref>. Comparatively, it is situated more towards the extracellular planes of the membrane bilayer <ref name="Article12"/>. The retinal is attached in the active site of rhodopsin through a protonated Schiff base (an N-substituted imine) bond to the ε-amino group of Lysine 296 residue (shown in <span style="color:lime;background-color:black;font-weight:bold;">green</span>) on the C-terminal Helix 7, with this linkage creating a positive charge on the chromophore <ref name="Article4"/>. The protonated Schiff base of rhodopsin is stabilized through <scene name=' | + | Rhodopsin consists of an opsin [http://en.wikipedia.org/wiki/Apoprotein apoprotein] and a <scene name='40/400594/Cv/7'>11-cis retinylidene chromophore</scene> in its active site. Rhodopsin is bound covalently to the 11-''cis'' retinal, the chromophore or "ligand," (shown in <span style="color:yellow;background-color:black;font-weight:bold;">yellow</span>) and this retinal is found in deeply in the core of the helices, in a hydrophobic site, parallel to the lipid bilayer<ref name="Article19">PMID:16051215</ref>. Comparatively, it is situated more towards the extracellular planes of the membrane bilayer <ref name="Article12"/>. The retinal is attached in the active site of rhodopsin through a protonated Schiff base (an N-substituted imine) bond to the ε-amino group of Lysine 296 residue (shown in <span style="color:lime;background-color:black;font-weight:bold;">green</span>) on the C-terminal Helix 7, with this linkage creating a positive charge on the chromophore <ref name="Article4"/>. The protonated Schiff base of rhodopsin is stabilized through <scene name='40/400594/Cv/8'>Glutamine 113</scene> residue electrostatic interaction with the counterion, holding the inactive rhodopsin in its state<ref name="Article20"/>. |
As this ligand is bound in the 12-s-''trans'' conformation, there arises the non-bonding interactions between the C-13 methyl group and C-10 hydrogen that contribute to non-planarity. This leads to the ability of the chromophore polyene tail to undergo fast photoisomerization around the C-11=C-12 double bond during light-induced activation<ref name="Article2">PMID:16962138</ref>. Also, it is found that the C-11=C-12 double bond is pre-twisted in the ground state of rhodopsin, which is partly attributed to the C20 methyl group attached to C13 through interaction with Tryptophan 265. This pre-twist may give insight on the features of isomerization about this bond upon light activation<ref name="ReferenceArticle"/>. | As this ligand is bound in the 12-s-''trans'' conformation, there arises the non-bonding interactions between the C-13 methyl group and C-10 hydrogen that contribute to non-planarity. This leads to the ability of the chromophore polyene tail to undergo fast photoisomerization around the C-11=C-12 double bond during light-induced activation<ref name="Article2">PMID:16962138</ref>. Also, it is found that the C-11=C-12 double bond is pre-twisted in the ground state of rhodopsin, which is partly attributed to the C20 methyl group attached to C13 through interaction with Tryptophan 265. This pre-twist may give insight on the features of isomerization about this bond upon light activation<ref name="ReferenceArticle"/>. | ||
Revision as of 09:38, 10 May 2017
| |||||||||||
3D structures of rhodopsin
Updated on 10-May-2017
References
- ↑ Hornak V, Ahuja S, Eilers M, Goncalves JA, Sheves M, Reeves PJ, Smith SO. Light activation of rhodopsin: insights from molecular dynamics simulations guided by solid-state NMR distance restraints. J Mol Biol. 2010 Feb 26;396(3):510-27. Epub 2009 Dec 11. PMID:20004206 doi:10.1016/j.jmb.2009.12.003
- ↑ 2.0 2.1 2.2 2.3 2.4 2.5 Sakmar TP. Structure of rhodopsin and the superfamily of seven-helical receptors: the same and not the same. Curr Opin Cell Biol. 2002 Apr;14(2):189-95. PMID:11891118
- ↑ 3.0 3.1 3.2 3.3 Kristiansen K. Molecular mechanisms of ligand binding, signaling, and regulation within the superfamily of G-protein-coupled receptors: molecular modeling and mutagenesis approaches to receptor structure and function. Pharmacol Ther. 2004 Jul;103(1):21-80. PMID:15251227 doi:10.1016/j.pharmthera.2004.05.002
- ↑ Millar RP, Newton CL. The year in G protein-coupled receptor research. Mol Endocrinol. 2010 Jan;24(1):261-74. Epub 2009 Dec 17. PMID:20019124 doi:10.1210/me.2009-0473
- ↑ 5.0 5.1 5.2 Meng EC, Bourne HR. Receptor activation: what does the rhodopsin structure tell us? Trends Pharmacol Sci. 2001 Nov;22(11):587-93. PMID:11698103
- ↑ 6.0 6.1 Shieh T, Han M, Sakmar TP, Smith SO. The steric trigger in rhodopsin activation. J Mol Biol. 1997 Jun 13;269(3):373-84. PMID:9199406 doi:10.1006/jmbi.1997.1035
- ↑ 7.0 7.1 7.2 7.3 7.4 7.5 7.6 7.7 7.8 Okada T, Ernst OP, Palczewski K, Hofmann KP. Activation of rhodopsin: new insights from structural and biochemical studies. Trends Biochem Sci. 2001 May;26(5):318-24. PMID:11343925
- ↑ 8.0 8.1 Okada T, Sugihara M, Bondar AN, Elstner M, Entel P, Buss V. The retinal conformation and its environment in rhodopsin in light of a new 2.2 A crystal structure. J Mol Biol. 2004 Sep 10;342(2):571-83. PMID:15327956 doi:10.1016/j.jmb.2004.07.044
- ↑ 9.0 9.1 Janz JM, Farrens DL. Assessing structural elements that influence Schiff base stability: mutants E113Q and D190N destabilize rhodopsin through different mechanisms. Vision Res. 2003 Dec;43(28):2991-3002. PMID:14611935
- ↑ 10.0 10.1 10.2 Kisselev OG. Focus on molecules: rhodopsin. Exp Eye Res. 2005 Oct;81(4):366-7. PMID:16051215 doi:10.1016/j.exer.2005.06.018
- ↑ 11.0 11.1 11.2 Verhoeven MA, Bovee-Geurts PH, de Groot HJ, Lugtenburg J, DeGrip WJ. Methyl substituents at the 11 or 12 position of retinal profoundly and differentially affect photochemistry and signalling activity of rhodopsin. J Mol Biol. 2006 Oct 13;363(1):98-113. Epub 2006 Jul 28. PMID:16962138 doi:10.1016/j.jmb.2006.07.039
- ↑ 12.0 12.1 12.2 12.3 Morris MB, Dastmalchi S, Church WB. Rhodopsin: structure, signal transduction and oligomerisation. Int J Biochem Cell Biol. 2009 Apr;41(4):721-4. Epub 2008 Aug 3. PMID:18692154 doi:10.1016/j.biocel.2008.04.025
- ↑ 13.0 13.1 13.2 13.3 13.4 Nelson, D., and Cox, M. Lehninger Principles of Biochemistry. 2008. 5th edition. W. H. Freeman and Company, New York, New York, USA. pp. 462-465.
- ↑ Hurley JB, Spencer M, Niemi GA. Rhodopsin phosphorylation and its role in photoreceptor function. Vision Res. 1998 May;38(10):1341-52. PMID:9667002
- ↑ 15.0 15.1 15.2 Park JH, Scheerer P, Hofmann KP, Choe HW, Ernst OP. Crystal structure of the ligand-free G-protein-coupled receptor opsin. Nature. 2008 Jul 10;454(7201):183-7. Epub 2008 Jun 18. PMID:18563085 doi:10.1038/nature07063
- ↑ 16.0 16.1 Surya A, Knox BE. Enhancement of opsin activity by all-trans-retinal. Exp Eye Res. 1998 May;66(5):599-603. PMID:9628807 doi:10.1006/exer.1997.0453
See Also
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, Wayne Decatur, Jaime Prilusky, Joel L. Sussman, Cinting Lim