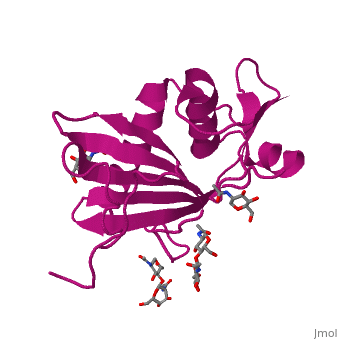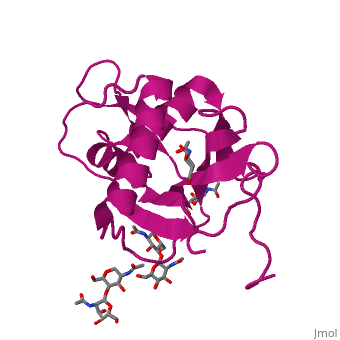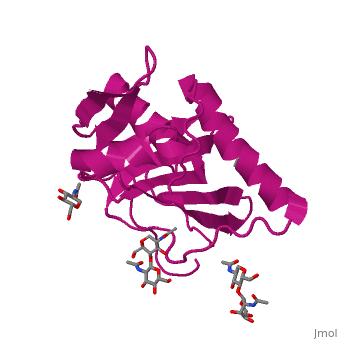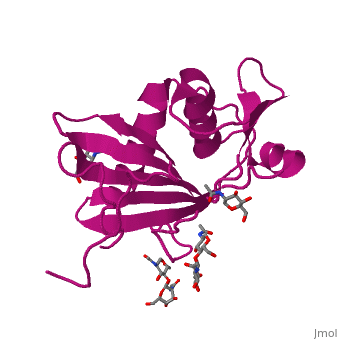This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
GP1 of Lassa Virus
From Proteopedia
(Difference between revisions)
| Line 7: | Line 7: | ||
GP1 of LASV is a 4 chain structure with <scene name='76/761695/Nag/1'>NAG</scene> ligands attached to each chain. The overall architecture of GP1 features a central β-sheet and two distinct halves: a glycosylated half containing the receptor-binding site that is made mostly by the central β-sheet and surrounding loops and a half that contains mostly helices and most likely faces the trimer axis<ref name="MolMechforLAMP1">DOI 10.1128/JVI.00651-15</ref>. The method used to determine this structure was [https://en.wikipedia.org/wiki/X-ray_crystallography X-ray diffraction] | GP1 of LASV is a 4 chain structure with <scene name='76/761695/Nag/1'>NAG</scene> ligands attached to each chain. The overall architecture of GP1 features a central β-sheet and two distinct halves: a glycosylated half containing the receptor-binding site that is made mostly by the central β-sheet and surrounding loops and a half that contains mostly helices and most likely faces the trimer axis<ref name="MolMechforLAMP1">DOI 10.1128/JVI.00651-15</ref>. The method used to determine this structure was [https://en.wikipedia.org/wiki/X-ray_crystallography X-ray diffraction] | ||
=== Histidine Triad=== | === Histidine Triad=== | ||
| - | Attached to this structure is a unique <scene name='76/761695/Histriad/ | + | Attached to this structure is a unique <scene name='76/761695/Histriad/6'>triad of histidines</scene> that is highly conserved among Old World arenaviruses. Located on the β-sheet face of GP1, the histidine triad is a structural element that directly interacts with [[LAMP1]] (Lysosome-associated membrane glycoprotein) and helps stabilize a LAMP1-"compatible" conformation by providing a molecular mechanism for the pH-dependent receptor switching<ref name="MolMechforLAMP1" />. The histidine triad is critical in forming a <scene name='76/761695/Lamp1bindingsite/3'>binding site</scene> for LAMP1. |
===LAMP1 Binding Site=== | ===LAMP1 Binding Site=== | ||
The primary cellular receptor of LASV α-dystroglycan (α-DG), which is recognized by a trimeric class 1 viral GPC on the surface of the virus. Following successful attachment to α-DG on cells, LASV is internalized via [https://en.wikipedia.org/wiki/Pinocytosis macropinocytosis], and the GPC facilitates membrane fusion at the acidic environment of a late endosomal compartment. Recent studies have shown that successful infection by LASV requires it to switch in a pH-dependent manner from α-DG to LAMP1. | The primary cellular receptor of LASV α-dystroglycan (α-DG), which is recognized by a trimeric class 1 viral GPC on the surface of the virus. Following successful attachment to α-DG on cells, LASV is internalized via [https://en.wikipedia.org/wiki/Pinocytosis macropinocytosis], and the GPC facilitates membrane fusion at the acidic environment of a late endosomal compartment. Recent studies have shown that successful infection by LASV requires it to switch in a pH-dependent manner from α-DG to LAMP1. | ||
Revision as of 10:10, 29 June 2017
| |||||||||||
References
- ↑ 1.0 1.1 Cohen-Dvashi H, Cohen N, Israeli H, Diskin R. Molecular mechanism for LAMP1 recognition by Lassa Virus. J Virol. 2015 May 13. pii: JVI.00651-15. PMID:25972533 doi:http://dx.doi.org/10.1128/JVI.00651-15
Categories: Cohen, N | Cohen-Dvashi, H | Diskin, R | Israeli, H | Arenavirus | Glycoprotein | Lassa | LASV | Receptor binding | Viral protein | 4zjf | GP1