This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Peptidase T
From Proteopedia
(Difference between revisions)
| Line 2: | Line 2: | ||
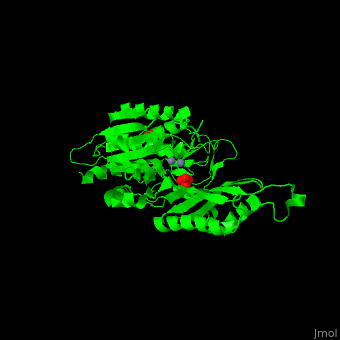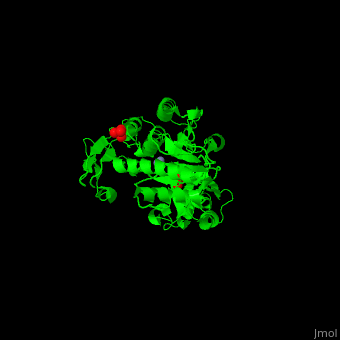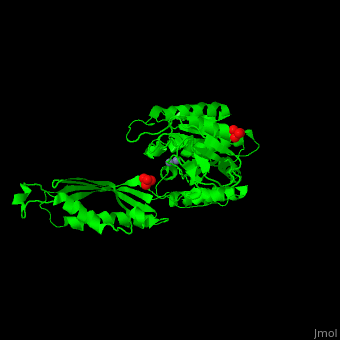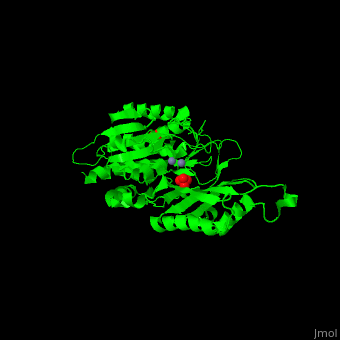
| - | '''Peptidase T''' (PT) removes the N-terminal peptide of tripeptides. PT is produced by bacteria. The tripeptides contain an N-terminal methionine, leucine or phenylalanine<ref>PMID:11856302</ref>. | + | '''Peptidase T''' (PT) removes the N-terminal peptide of tripeptides. PT is produced by bacteria. The tripeptides contain an N-terminal methionine, leucine or phenylalanine<ref>PMID:11856302</ref>. |
| + | |||
| + | *<scene name='50/501403/Cv/2'>Zn coordinations sites</scene>. Water molecule shown as red sphere. | ||
</StructureSection> | </StructureSection> | ||
==3D structures of peptidase T== | ==3D structures of peptidase T== | ||
Revision as of 09:37, 28 August 2017
| |||||||||||
3D structures of peptidase T
Updated on 28-August-2017
1fno – PT – Salmonella typhymurium
3gb0 – PT – Bacillus cereus
3ife - PT – Bacillus anthracis
1vix – PT – Escherichia coli