This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Prolyl hydroxylase domain
From Proteopedia
| Line 1: | Line 1: | ||
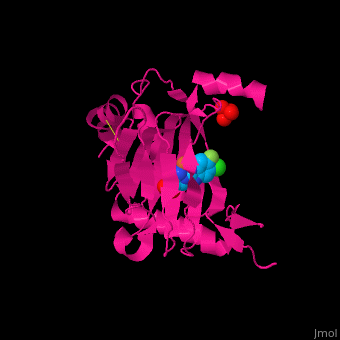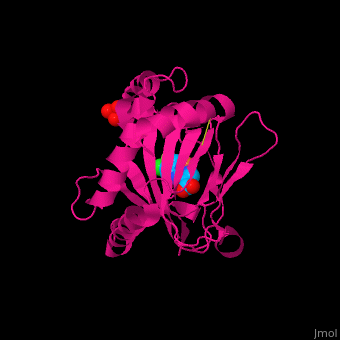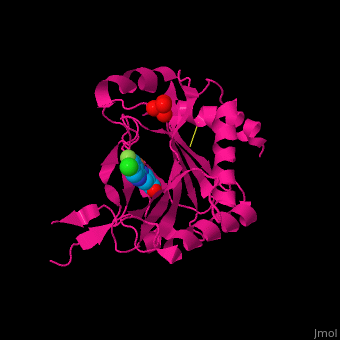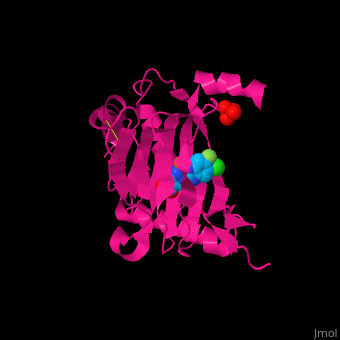
| - | <StructureSection load='3ouh' size='400' side='right' scene='' caption='Human PHD2 catalytic domain complex with Fe+2 ion (orange), inhibitor and sulfate, [[3ouh]]'> | + | <StructureSection load='3ouh' size='400' side='right' scene='45/459221/Cv/1' caption='Human PHD2 catalytic domain complex with Fe+2 ion (orange), inhibitor and sulfate, [[3ouh]]'> |
'''Prolyl hydroxylase domain''' (PHD) proteins mediate oxygen-dependent degradation of Hypoxia-inducible factor (HIF) α subunit. They include PHD1, PHD2 and PHD3. The PHD is a Fe+2/oxogluterate (2OG)-dependent enzyme. [[3ouh]] is the crystallized structure of the enzyme PHD2, an [[oxidoreductase]] that is 237 amino acids long with a molecular weight of 27 kDa. [[3ouh]] is found in [http://en.wikipedia.org/wiki/Homo_sapiens Homo sapiens] and is a homolog of [http://en.wikipedia.org/wiki/EGLN1 EGLN1] found in [http://en.wikipedia.org/wiki/Caenorhabditis_elegans C. elegans]. | '''Prolyl hydroxylase domain''' (PHD) proteins mediate oxygen-dependent degradation of Hypoxia-inducible factor (HIF) α subunit. They include PHD1, PHD2 and PHD3. The PHD is a Fe+2/oxogluterate (2OG)-dependent enzyme. [[3ouh]] is the crystallized structure of the enzyme PHD2, an [[oxidoreductase]] that is 237 amino acids long with a molecular weight of 27 kDa. [[3ouh]] is found in [http://en.wikipedia.org/wiki/Homo_sapiens Homo sapiens] and is a homolog of [http://en.wikipedia.org/wiki/EGLN1 EGLN1] found in [http://en.wikipedia.org/wiki/Caenorhabditis_elegans C. elegans]. | ||
Revision as of 10:29, 10 September 2017
| |||||||||||
3D Structures of prolyl hydroxylase domain
2y33 – hPHD2 catalytic domain + Zn + chloro-iodoisoquinolin-carbonyl glycine – human
2y34 - hPHD2 catalytic domain + Fe + chloro-iodoisoquinolin-carbonyl glycine
3ouh, 3oui - hPHD2 catalytic domain + Fe + inhibitor
3ouj - hPHD2 catalytic domain + Fe + 2OG
3hqr - hPHD2 catalytic domain (mutant) + Mn + HIF 1 α C terminal + oxalylglycine
3hqu - hPHD2 catalytic domain + Fe + HIF 1 α C terminal + chloro-iodoisoquinolin-carbonyl glycine
2hbt, 2hbu - hPHD2 catalytic domain + Fe + chloro-iodoisoquinolin-carbonyl glycine
2g19, 2g1m - hPHD2 catalytic domain + Fe + hydroxy-iodoisoquinolin-carbonyl glycine
4h6j – hPHD Pasb domain (mutant) + aryl hydrocarbon nuclear translocator (mutant)
References
- ↑ Stolze IP, Mole DR, Ratcliffe PJ. Regulation of HIF: prolyl hydroxylases. Novartis Found Symp. 2006;272:15-25; discussion 25-36. PMID:16686427
- ↑ Rosen M D, Venkatesan H, Peltier H M, Bembenek S D, Kanelakis K C, Zhao L X, Leonard B E, Hocutt F M, Wu X, Palomino H L, Brondtetter T I, Haugh P V, Cagnon L, Yan W, Liotta L A, Young A, Mirzadegan T, Shankley N P, Barrett T D, Rabinowitz M H. Benzimidazole-2-pyrazole HIF Prolyl 4-Hydroxylase Inhibitors as Oral Erythropoietin Secretagogues. ACS Medicinal Chemical Letters. 2010 Oct 5.
Created with the participation of Andrew Winslow.