This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Retroviral Integrase
From Proteopedia
| Line 3: | Line 3: | ||
==Function== | ==Function== | ||
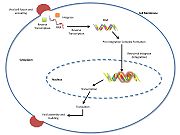
| - | [[Retroviral Integrase]] is an essential retroviral enzyme that binds to viral DNA and inserts it into a host cell chromosome. The reverse transcribed cDNA of human immunodeficiency virus type 1 (HIV-1) is inserted in the host cell genome in order increase pathogen fitness and virulence. Integrase is produced by a class of retrovirus (like HIV) and is used by the virus to incorporate its genetic material into the host cell DNA. The host cellular machinery then produces mRNA and then protein from the incorporated genetic material, thus replicating the virus. Although several integrase inhibiting drugs have been investigated, the mechanism responsible for strand-transfer inhibition action remains to be elucidated. However, | + | [[Retroviral Integrase]] is an essential retroviral enzyme that binds to viral DNA and inserts it into a host cell chromosome. The reverse transcribed cDNA of human immunodeficiency virus type 1 (HIV-1) is inserted in the host cell genome in order increase pathogen fitness and virulence. Integrase is produced by a class of retrovirus (like HIV) and is used by the virus to incorporate its genetic material into the host cell DNA. The host cellular machinery then produces mRNA and then protein from the incorporated genetic material, thus replicating the virus. Although several integrase inhibiting drugs have been investigated, the mechanism responsible for strand-transfer inhibition action remains to be elucidated. However, Hare el al (2010)<ref name=Hare_2010>PMID:20118915</ref> determined the structural constituents of retroviral integration. Further elucidation of the complete structure of the retroviral integrase, and its application to regulate functional and enzymatic activities could potentially enable researchers to delay the progression of retroviral diseases. Moreover, study of HIV-1 integration could lead to a promising new target, and contribute to the generation pharmacophore models for antiviral therapy. <br/> |
HIV Integrase inhibitors: Raltegravir, marketed as Isentress is currently approved as a therapeutic inhibitor of HIV integrase. It was approved on October 12, 2007. | HIV Integrase inhibitors: Raltegravir, marketed as Isentress is currently approved as a therapeutic inhibitor of HIV integrase. It was approved on October 12, 2007. | ||
[See below for a table of antiretroviral drugs with trade name, company, patents, and notes.] | [See below for a table of antiretroviral drugs with trade name, company, patents, and notes.] | ||
| Line 30: | Line 30: | ||
==PFV Intasome Crystallization== | ==PFV Intasome Crystallization== | ||
| - | To mimic the viral DNA ends of HIV-1, Hare ''et al'' (2010) utilized soluble and fully functional prototype foamy virus (PFV) intasome preparations, obtained using recombinant PFV integrase and double-stranded oligonucleotides. | + | To mimic the viral DNA ends of HIV-1, Hare ''et al'' (2010)<ref name="Hare_2010" /> utilized soluble and fully functional prototype foamy virus (PFV) intasome preparations, obtained using recombinant PFV integrase and double-stranded oligonucleotides. |
The remarkable stability of the integrase-DNA complexes were determined by observing the ''in vitro'' strand transfer reactions, which were classified into three modes of deproteination migration: (1) single concerted events: linearized target plasmid; (2) multiple concerted events: smear; (3) half-site events: open circular DNA. Further characterization of the PFV intasome also exhibited structural substantiality which implied strong protein-protein and protein-DNA interactions despite prolonged incubation under high ionic strength conditions. Comprehensive crystallization assays effected a viable crystal configuration that diffracted X-rays to 2.9 Angstroms resolution. A three-dimensional structure was ultimately determined. The asymmetric unit contained a single integrase dimer with a stably bound viral DNA molecule, and a pair of integrase dimers consociated with symmetry, which formed an oblong tetramer. The dimer interface is stabilized by intermolecular amino terminal and catalytic core domains (inner subunit-outer subunit) interactions. The overall shape of the oblong tetramer is unique albeit bearing semblances to previously reported HIV-1 integrase complexes. | The remarkable stability of the integrase-DNA complexes were determined by observing the ''in vitro'' strand transfer reactions, which were classified into three modes of deproteination migration: (1) single concerted events: linearized target plasmid; (2) multiple concerted events: smear; (3) half-site events: open circular DNA. Further characterization of the PFV intasome also exhibited structural substantiality which implied strong protein-protein and protein-DNA interactions despite prolonged incubation under high ionic strength conditions. Comprehensive crystallization assays effected a viable crystal configuration that diffracted X-rays to 2.9 Angstroms resolution. A three-dimensional structure was ultimately determined. The asymmetric unit contained a single integrase dimer with a stably bound viral DNA molecule, and a pair of integrase dimers consociated with symmetry, which formed an oblong tetramer. The dimer interface is stabilized by intermolecular amino terminal and catalytic core domains (inner subunit-outer subunit) interactions. The overall shape of the oblong tetramer is unique albeit bearing semblances to previously reported HIV-1 integrase complexes. | ||
| Line 42: | Line 42: | ||
===Crystallographic and Refinement Statistics=== | ===Crystallographic and Refinement Statistics=== | ||
| - | Hare ''et al'' (2010) have published data on seven crystal structures. These data include the PFV IN complex (apo form) and six additional structures, including the complex bound to Mg, Mn, Mg/MK0518, Mn/MK0518, Mg/GS9137, and Mn/GS9137. All seven structures belong to the P41212 space group. They have been refined to between 2.85 and 3.25 Å resolution. | + | Hare ''et al'' (2010)<ref name="Hare_2010" /> have published data on seven crystal structures. These data include the PFV IN complex (apo form) and six additional structures, including the complex bound to Mg, Mn, Mg/MK0518, Mn/MK0518, Mg/GS9137, and Mn/GS9137. All seven structures belong to the P41212 space group. They have been refined to between 2.85 and 3.25 Å resolution. |
==Overall Architecture & Components== | ==Overall Architecture & Components== | ||
| Line 48: | Line 48: | ||
===Structure=== | ===Structure=== | ||
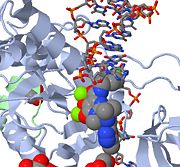
| - | The overall structure of the assembled PFV intasome is a tetramer model based on two domain structures with a dimer-dimer interface. Previous intasome models depict a similar but more flexible structure while the PFV intasome has been shown to be highly constrained. Using homology modeling, Hare ''et al'' (2010) propose that shorter interdomain linkers may be a factor in flexibility, specifically in HIV-1 integrase. The inner subunits of the tetramer are implicated in the overall tetramerization and viral DNA binding. The catalytic core domains of the outer subunits may act as supports, but since the amino- and carboxy-terminal domains are unresolved in electron density maps, their function remains inconclusive. The catalytic core domain and carboxy-terminal domain linker adopts an extended conformation for most of its length, and are located parallel to the amino-terminal domain and catalytic core domain linker of the inner subunit. The interdomain linkers The interdomain linkers (CCD-CTD linker and NTD-CCD linker) bind both halves of the intasome together, and the structure is further stabilized by a pair of carboxy-terminal domains interacting with both inner catalytic core domains. | + | The overall structure of the assembled PFV intasome is a tetramer model based on two domain structures with a dimer-dimer interface. Previous intasome models depict a similar but more flexible structure while the PFV intasome has been shown to be highly constrained. Using homology modeling, Hare ''et al'' (2010)<ref name="Hare_2010" /> propose that shorter interdomain linkers may be a factor in flexibility, specifically in HIV-1 integrase. The inner subunits of the tetramer are implicated in the overall tetramerization and viral DNA binding. The catalytic core domains of the outer subunits may act as supports, but since the amino- and carboxy-terminal domains are unresolved in electron density maps, their function remains inconclusive. The catalytic core domain and carboxy-terminal domain linker adopts an extended conformation for most of its length, and are located parallel to the amino-terminal domain and catalytic core domain linker of the inner subunit. The interdomain linkers The interdomain linkers (CCD-CTD linker and NTD-CCD linker) bind both halves of the intasome together, and the structure is further stabilized by a pair of carboxy-terminal domains interacting with both inner catalytic core domains. |
===Integrase and DNA interactions=== | ===Integrase and DNA interactions=== | ||
Revision as of 15:59, 27 September 2017
| |||||||||||
Integrase Inhibitors
| Name | Brand | Company | Patent | Notes |
| Raltegravir | Isentress | Merck & Co. | - | also known as MK-0518. The isopropyl and methyl-oxadiazole of MK-0518 are involved in hydrophobic and stacking interactions with side chains of Pro 214 and Tyr 212 to stabilize this drug within the PFV intasome active site. This manner of drug-binding interaction causes displacement of the reactive 3' viral DNA end from the active site of PFV intasome. After binding of MK-0518 to active site, the reactive 3' hydroxyl group moves away from the active site of the PFV intasome by more than 6 Angstroms. Raltegravir was approved by the FDA on October 12, 2007, for use with other anti-HIV agents in the treatment of HIV infection in adults. It is the first integrase inhibitor approved by the FDA. |
| Elvitegravir | - | Gilead Science | - | GS-9137 interacts with Pro 214 of PFV intasome through its quinolone base and isopropyl group. In experimental stages; shares the core structure of quinolone antibiotics. Phase II studies of elvitegravir in people who are treatment experienced have been completed. Phase III studies in treatment experienced patients are ongoing. A phase II study of elvitegravir in people who have never taken antiretroviral therapy is underway. This study will also be evaluated a boosting agent in place of Norvir, currently called GS9350. Elvitegravir holds promise for HIV-positive patients who have taken other anti-HIV drugs in the past. |
| MK-2048 | - | Merck & Co. | - | A second generation integrase inhibitor, intended to be used against HIV infection. It is superior to the first available integrase inhibitor, raltegravir, in that it inhibits the HIV enzyme integrase 4 times longer. It is being investigated for use as part of pre-exposure prophylaxis (PrEP). |
See also Retroviral Integrase Inhibitor Pharmacokinetics[2].
Additional Resources
For additional information, see: Human Immunodeficiency Virus
3D Structures of Retroviral Integrase
Updated on 27-September-2017
References
- ↑ 1.0 1.1 1.2 1.3 Hare S, Gupta SS, Valkov E, Engelman A, Cherepanov P. Retroviral intasome assembly and inhibition of DNA strand transfer. Nature. 2010 Mar 11;464(7286):232-6. Epub 2010 Jan 31. PMID:20118915 doi:10.1038/nature08784
- ↑ Iwamoto M, Wenning LA, Petry AS, Laethem M, De Smet M, Kost JT, Breidinger SA, Mangin EC, Azrolan N, Greenberg HE, Haazen W, Stone JA, Gottesdiener KM, Wagner JA. Minimal effects of ritonavir and efavirenz on the pharmacokinetics of raltegravir. Antimicrob Agents Chemother. 2008 Dec;52(12):4338-43. Epub 2008 Oct 6. PMID:18838589 doi:10.1128/AAC.01543-07
2. http://www.isentress.com/raltegravir/isentress/consumer/index.jsp
3. deJesus, Edwin HIV Antiretroviral Agents in Development. The Body: The Complete HIV/AIDS Resource. March 30, 2006.
4. AIDS Info
5. Krishan K. Pandey and Duane P. Grandgenett (2008) HIV-1 Integrase Strand Transfer Inhibitors: Novel Insights into their Mechanism of Action. Retrovirology: Research and Treatment" 2008:2 11-16
6.James F. Braun, DO, Ruth J. Cronje, PhD, Marnie G. Henderson (2008) HIV-1 Integrase Inhibitors. www.prn.org Volume 13, Pages 1–9
Further Reading
- GEN News Highlights "Scientists Solve 3-D Crystal Structure of Retroviral Integrase Bound to Viral DNA", Genetic Engineering & Biotechnology News February 1, 2010.
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Rhysly Martinez, Joel L. Sussman, Alexander Berchansky, David Canner, Jordan Heard, Eugene Babcock, Garrett Asanuma