This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Pyruvate phosphate dikinase
From Proteopedia
(Difference between revisions)
| Line 10: | Line 10: | ||
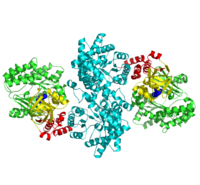
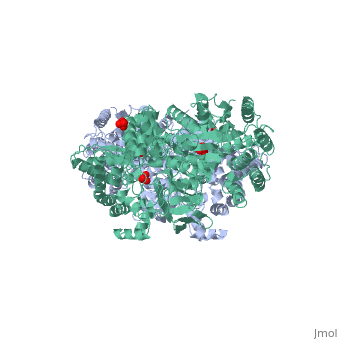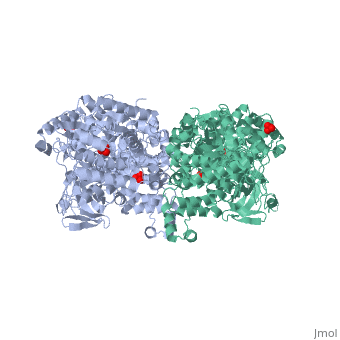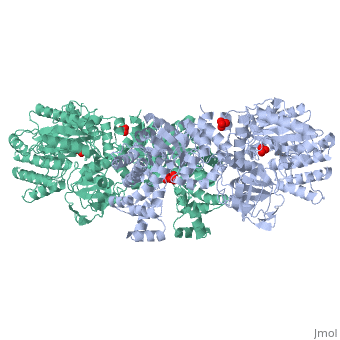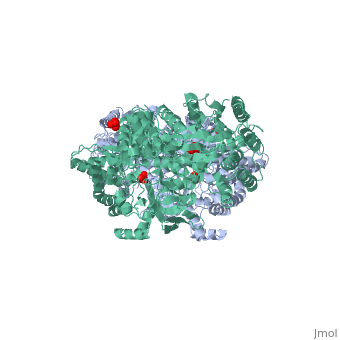
PPDK assembles into homodimers of ~95 kD subunit molecular mass. The monomer is comprised of three domains and contains two distinct reaction centers located ~45 Å apart; the PEP/pyruvate partial reaction (step 1) takes place at the C-terminal domain (adopting an α/β barrel fold) and the nucleotide and inorganic phosphate partial reactions (steps 2 and 3) take place at the N-terminal domain (adopting the ATP grasp fold with two sub domains). A central domain, tethered to the N- and C-terminal domains by two closely-associated linkers, contains a phosphorylatable histidine residue (His455). To shuttle the phosphoryl group between the two reaction centers, the His-domain undergoes domain motion of ~110° swivel around the two linkers. In addition, upon detachment from the His-domain, the two nucleotide-binding sub domains undergo a ~40° hinge motion that opens the active site cleft. | PPDK assembles into homodimers of ~95 kD subunit molecular mass. The monomer is comprised of three domains and contains two distinct reaction centers located ~45 Å apart; the PEP/pyruvate partial reaction (step 1) takes place at the C-terminal domain (adopting an α/β barrel fold) and the nucleotide and inorganic phosphate partial reactions (steps 2 and 3) take place at the N-terminal domain (adopting the ATP grasp fold with two sub domains). A central domain, tethered to the N- and C-terminal domains by two closely-associated linkers, contains a phosphorylatable histidine residue (His455). To shuttle the phosphoryl group between the two reaction centers, the His-domain undergoes domain motion of ~110° swivel around the two linkers. In addition, upon detachment from the His-domain, the two nucleotide-binding sub domains undergo a ~40° hinge motion that opens the active site cleft. | ||
| - | [[Image:1KBL dimer.png| | + | [[Image:1KBL dimer.png|center|thumb|'''PPDK dimer'''<br>PEP-binding domain - cyan; nucleotide binding domain - green; His-domain - yellow; domain linkers -red; phosphorylatable His455 - blue spheres|200px]] |
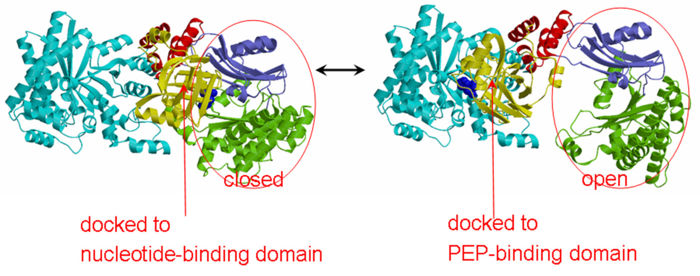
'''The His-domain in the two conformational states of PPDK. His455 is shown in blue spheres:'''[[Image:two_cond.jpg|right|700px]]<br><br><br> | '''The His-domain in the two conformational states of PPDK. His455 is shown in blue spheres:'''[[Image:two_cond.jpg|right|700px]]<br><br><br> | ||
Revision as of 08:04, 22 October 2017
Pyruvate Phosphate Dikinase - a Molecular Machine
| |||||||||||
3D Structures of PPDK
Updated on 22-October-2017
Proteopedia Page Contributors and Editors (what is this?)
Joel L. Sussman, Osnat Herzberg, Michal Harel, Jaime Prilusky, Alexander Berchansky, Dan Bolser, Karl Oberholser, David Canner, Eran Hodis