This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1053
From Proteopedia
(Difference between revisions)
| Line 5: | Line 5: | ||
===Operon Overview=== | ===Operon Overview=== | ||
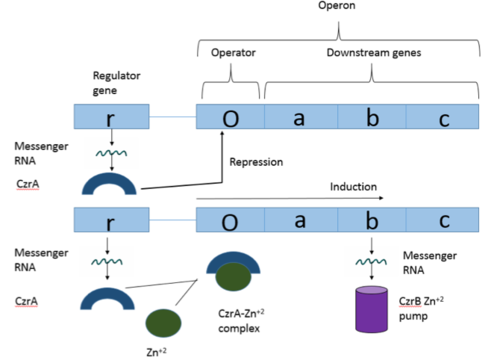
[https://en.wikipedia.org/wiki/Operon Operons] are a critical genetic component of most prokaryotic cells. There are many different operons, responsible for the production of proteins with a wide range of functions. The most well-known and studied operons are the [https://en.wikipedia.org/wiki/Lac_operon Lac] and [https://en.wikipedia.org/wiki/Trp_operon Trp] operons, responsible for producing enzymes which metabolize lactose and tryptophan respectively. Despite many differences in each operon and the proteins that they encode, operons all function in the same general manner (Figure 1). Each operon contains a [https://en.wikipedia.org/wiki/Regulator_gene regulator], an [https://en.wikipedia.org/wiki/Operator_(biology) operator], and one or more [https://en.wikipedia.org/wiki/Structural_gene structural genes]. The regulator gene codes for a protein responsible for managing the expression level of the structural genes. The operator contains the binding sequence for [https://en.wikipedia.org/wiki/RNA_polymerase RNA polymerase] and is the site where [https://en.wikipedia.org/wiki/Transcription_(biology) transcription] begins. Lastly, the structural genes code for proteins to be used elsewhere. The regulator protein (produced as a result of expression of the regulator gene) usually acts in a repressive manner. The regulator protein will bind to the operator gene, inhibiting the binding and/or progression of RNA polymerase to the structural genes, thus inhibiting transcription of the genes into mRNA. If the regulator protein were always active, the structural genes would never be expressed, so there must be a way to inactive the regulator protein, thus enabling expression of the structural genes. This is usually achieved through the binding of an inhibitor to the regulator protein. Since regulator proteins are DNA binding proteins, often this inhibition is [https://en.wikipedia.org/wiki/Allosteric_regulation allosteric] rather than competitive. The inhibitor of the regulator protein binds to somewhere other than the active site of the protein, changing the conformation of the regulator protein to decrease its ability to bind DNA and repress transcription. | [https://en.wikipedia.org/wiki/Operon Operons] are a critical genetic component of most prokaryotic cells. There are many different operons, responsible for the production of proteins with a wide range of functions. The most well-known and studied operons are the [https://en.wikipedia.org/wiki/Lac_operon Lac] and [https://en.wikipedia.org/wiki/Trp_operon Trp] operons, responsible for producing enzymes which metabolize lactose and tryptophan respectively. Despite many differences in each operon and the proteins that they encode, operons all function in the same general manner (Figure 1). Each operon contains a [https://en.wikipedia.org/wiki/Regulator_gene regulator], an [https://en.wikipedia.org/wiki/Operator_(biology) operator], and one or more [https://en.wikipedia.org/wiki/Structural_gene structural genes]. The regulator gene codes for a protein responsible for managing the expression level of the structural genes. The operator contains the binding sequence for [https://en.wikipedia.org/wiki/RNA_polymerase RNA polymerase] and is the site where [https://en.wikipedia.org/wiki/Transcription_(biology) transcription] begins. Lastly, the structural genes code for proteins to be used elsewhere. The regulator protein (produced as a result of expression of the regulator gene) usually acts in a repressive manner. The regulator protein will bind to the operator gene, inhibiting the binding and/or progression of RNA polymerase to the structural genes, thus inhibiting transcription of the genes into mRNA. If the regulator protein were always active, the structural genes would never be expressed, so there must be a way to inactive the regulator protein, thus enabling expression of the structural genes. This is usually achieved through the binding of an inhibitor to the regulator protein. Since regulator proteins are DNA binding proteins, often this inhibition is [https://en.wikipedia.org/wiki/Allosteric_regulation allosteric] rather than competitive. The inhibitor of the regulator protein binds to somewhere other than the active site of the protein, changing the conformation of the regulator protein to decrease its ability to bind DNA and repress transcription. | ||
| - | [[Image:Operon.png|500px|thumb|center|Figure 1: Overview of operon structure]] | + | [[Image:Operon.png|500px|thumb|center|Figure 1: Overview of CzrA operon structure]] |
===The Czr Operon=== | ===The Czr Operon=== | ||
The <u>C</u>hromosome determined <u>z</u>inc <u>r</u>esponsible (Czr) operon acts as described above (Figure 1), with CzrA acting as a regulator protein to the downstream structural gene CzrB<ref name="critical">Arunkumar A., Campanello G., Giedroc D. (2009). Solution Structure of a | The <u>C</u>hromosome determined <u>z</u>inc <u>r</u>esponsible (Czr) operon acts as described above (Figure 1), with CzrA acting as a regulator protein to the downstream structural gene CzrB<ref name="critical">Arunkumar A., Campanello G., Giedroc D. (2009). Solution Structure of a | ||
Revision as of 00:33, 19 January 2018
Zinc Dependent Transcriptional Repressor of the Czr operon (CzrA)
| |||||||||||
References
- ↑ 1.0 1.1 1.2 1.3 Arunkumar A., Campanello G., Giedroc D. (2009). Solution Structure of a paradigm ArsR family zinc sensor in the DNA-bound state. PNAS 106:43 18177-18182.
- ↑ MacPherson S, Larochelle M, Turcotte B. A fungal family of transcriptional regulators: the zinc cluster proteins. Microbiol Mol Biol Rev. 2006 Sep;70(3):583-604. PMID:16959962 doi:http://dx.doi.org/10.1128/MMBR.00015-06
- ↑ Miller J, McLachlan AD, Klug A. Repetitive zinc-binding domains in the protein transcription factor IIIA from Xenopus oocytes. EMBO J. 1985 Jun 4;4(6):1609-1614.
- ↑ Grossoehme NE, Giedroc DP. Energetics of allosteric negative coupling in the zinc sensor S. aureus CzrA. J Am Chem Soc. 2009 Dec 16;131(49):17860-70. doi: 10.1021/ja906131b. PMID:19995076 doi:http://dx.doi.org/10.1021/ja906131b