User:Neel Bhagat/Sandbox 1
From Proteopedia
| Line 1: | Line 1: | ||
== Introduction == | == Introduction == | ||
| - | + | ||
=== Overview === | === Overview === | ||
The SRp20 protein is an alternative splicing factor found in homo sapiens as well as many other eukaryotic organisms. It is a relatively small protein with a length of 164 amino acids and a weight of about 19kDa. In fact, it is the smallest member of the SR protein family. The protein contains two domains: a serine-arginine rich (SR) domain and a RNA-recognition domain (RRM). | The SRp20 protein is an alternative splicing factor found in homo sapiens as well as many other eukaryotic organisms. It is a relatively small protein with a length of 164 amino acids and a weight of about 19kDa. In fact, it is the smallest member of the SR protein family. The protein contains two domains: a serine-arginine rich (SR) domain and a RNA-recognition domain (RRM). | ||
| Line 9: | Line 9: | ||
=== Structure Determination === | === Structure Determination === | ||
| + | <StructureSection load='2i2y' size='400' side='right' caption='SRp20 bound to RNA ligand and IgG binding domain 1 (PDB entry [[2i2y]])' scene=''> | ||
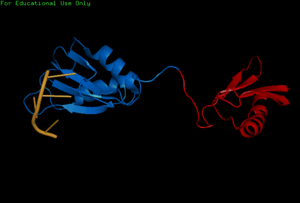
[[Image:BothChains.png|300px|right|thumb|'''Figure 1.''' This is a figure legend]] | [[Image:BothChains.png|300px|right|thumb|'''Figure 1.''' This is a figure legend]] | ||
Attempts to determine the structure of native SRP20 were largely unsuccessful due to the low solubility of the protein. This is likely due to the hydrophobic core of the RRM and exposed hydrophobic residues for RNA recognition on the β-sheets. As a solution, researchers removed the SR domain from the C terminus, leaving only the SRP20 RRM and a small arginine rich segment at the C terminus, then fused with a soluble <scene name='78/782597/Imager0/2'>IgG binding domain</scene> of Streptococcal protein G to the N terminus of the protein, providing the first published structure of the SRP20 RRM via NMR3. However, the solution of the structure via NMR, in addition to fusion with a globular tag, results in multiple possible conformations of the protein, meaning measurements such as bond angles, lengths, and substrate interactions are variable. Further, information concerning structural aspects of the SR domain are still limited to experimental data of protein function with certain mutations or deletions, and by comparison to sister proteins such as 9G8. To date, structure of the SR domain or the protein without the globular tag have not been solved, nor has a crystal structure for any part of the protein been determined. | Attempts to determine the structure of native SRP20 were largely unsuccessful due to the low solubility of the protein. This is likely due to the hydrophobic core of the RRM and exposed hydrophobic residues for RNA recognition on the β-sheets. As a solution, researchers removed the SR domain from the C terminus, leaving only the SRP20 RRM and a small arginine rich segment at the C terminus, then fused with a soluble <scene name='78/782597/Imager0/2'>IgG binding domain</scene> of Streptococcal protein G to the N terminus of the protein, providing the first published structure of the SRP20 RRM via NMR3. However, the solution of the structure via NMR, in addition to fusion with a globular tag, results in multiple possible conformations of the protein, meaning measurements such as bond angles, lengths, and substrate interactions are variable. Further, information concerning structural aspects of the SR domain are still limited to experimental data of protein function with certain mutations or deletions, and by comparison to sister proteins such as 9G8. To date, structure of the SR domain or the protein without the globular tag have not been solved, nor has a crystal structure for any part of the protein been determined. | ||
Revision as of 00:45, 31 March 2018
Contents |
Introduction
Overview
The SRp20 protein is an alternative splicing factor found in homo sapiens as well as many other eukaryotic organisms. It is a relatively small protein with a length of 164 amino acids and a weight of about 19kDa. In fact, it is the smallest member of the SR protein family. The protein contains two domains: a serine-arginine rich (SR) domain and a RNA-recognition domain (RRM).
History
Splicing is one step in the process of RNA maturation that cuts out introns and joins exons together. Both the spliceosome, a complex of snRNAs (U1, U2, etc.), and splicing factors like SRp20 interact with intron consensus sequences in the pre-mRNA to regulate this process. Alternative splicing allows one mRNA molecule to produce numerous proteins that perform different functions in a cell by inclusion and exclusion of RNA sequences. There are two main families of splicing factors: Serine-Arginine rich (SR) proteins and heterogeneous nuclear RiboNucleoProteins (hnRNPs). The SRp20 protein belongs to the SR protein family. All SR proteins are defined by a RNA-binding domain at the N-terminus and a serine-arginine rich domain at the C-terminus (Corbo et al. 2013). The discovery of this family started in the 1900s with the SF2 (SRp30a) protein and has since come to include twelve proteins, all of which act as splicing factors. SRp20 was first discovered in calf thymus when it was separated with several other SR proteins based on their molecular weight (Zhaler 1992). An identical protein, X16, was discovered in an earlier paper studying different genes that change expression during B-cell development (Ayane 1991). At the time, the protein was assumed to play a role in RNA processing and cellular proliferation, a finding that was later proved to be true (Ayane 1991; Corbo et al. 2013).
Structure Determination
| |||||||||||