This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Isabella Gieck/Sandbox 1
From Proteopedia
| Line 3: | Line 3: | ||
== Introduction == | == Introduction == | ||
Hrp1 is a heterogeneous ribonuclear protein of Saccharomyces cerevisiae, baker’s yeast. Hrp1 is an essential component of 3’ pre-mRNA processing and contributes to the preparatory cleavage required for polyadenylation. The gene expressed as Hrp1, HRP1, was first isolated by Henry, et al.<ref> Henry, Michael, et al. “Potential RNA Binding Proteins in Saccharomyces Cerevisiae Identified as Suppressors of Temperature-Sensitive Mutations inNPL3.” Genetics, vol. 142, Jan. 1996, pp. 103–115. </ref> and was later attributed to the Hrp1 protein by Kessler, et al.<ref> Kessler, Marco M, et al. “Purification of the Saccharomyces Cerevisiae Cleavage/Polyadenylation Factor I.” Journal of Biological Chemistry, vol. 271, no. 43, 25 Oct. 1996, pp. 27167–27175. </ref> Hrp1 also participates in the regulation of the 3’ end. | Hrp1 is a heterogeneous ribonuclear protein of Saccharomyces cerevisiae, baker’s yeast. Hrp1 is an essential component of 3’ pre-mRNA processing and contributes to the preparatory cleavage required for polyadenylation. The gene expressed as Hrp1, HRP1, was first isolated by Henry, et al.<ref> Henry, Michael, et al. “Potential RNA Binding Proteins in Saccharomyces Cerevisiae Identified as Suppressors of Temperature-Sensitive Mutations inNPL3.” Genetics, vol. 142, Jan. 1996, pp. 103–115. </ref> and was later attributed to the Hrp1 protein by Kessler, et al.<ref> Kessler, Marco M, et al. “Purification of the Saccharomyces Cerevisiae Cleavage/Polyadenylation Factor I.” Journal of Biological Chemistry, vol. 271, no. 43, 25 Oct. 1996, pp. 27167–27175. </ref> Hrp1 also participates in the regulation of the 3’ end. | ||
| + | |||
| + | <StructureSection load='2cjk' size='400' side='right' caption='(PDB entry [[2cjk]])' scene=''> | ||
== Function == | == Function == | ||
| Line 20: | Line 22: | ||
Hrp1 participates in multiple regulatory pathways involving 3’ processing.<ref> Tuck, Alex C, and David Tollervey. “A Transcriptome-Wide Atlas of RNP Composition Reveals Diverse Classes of MRNAs and LncRNAs.” Cell, vol. 154, 29 Aug. 2013, pp. 996–1009. </ref> It has been implicated in the inhibition of transcription by the Sen-1 helicase as mediated by Nrd-1.<ref> Kuehner, Jason N., and David A. Brow. “Regulation of a Eukaryotic Gene by GTP-Dependent Start Site Selection and Transcription Attenuation.” Molecular Cell, vol. 31, no. 2, 2008, pp. 201–211., doi:10.1016/j.molcel.2008.05.018. </ref> Hrp1 also activates the nonsense-mediated mRNA decay pathway, which monitors translation and degrades incorrect mRNA.<ref> González, Carlos I., et al. “The Yeast HnRNP-like Protein Hrp1/Nab4 Marks a Transcript for Nonsense-Mediated MRNA Decay.” Molecular Cell, vol. 5, no. 3, 2000, pp. 489–499., doi:10.1016/s1097-2765(00)80443-8. </ref> Hrp1, along with Rna14, competes with Npl13, which inhibits cleavage at the polyadenylation site.<ref> Bucheli, M. E., et al. “Polyadenylation Site Choice in Yeast Is Affected by Competition between Npl3 and Polyadenylation Factor CFI.” RNA, vol. 13, no. 10, 2007, pp. 1756–1764., doi:10.1261/rna.607207. </ref> | Hrp1 participates in multiple regulatory pathways involving 3’ processing.<ref> Tuck, Alex C, and David Tollervey. “A Transcriptome-Wide Atlas of RNP Composition Reveals Diverse Classes of MRNAs and LncRNAs.” Cell, vol. 154, 29 Aug. 2013, pp. 996–1009. </ref> It has been implicated in the inhibition of transcription by the Sen-1 helicase as mediated by Nrd-1.<ref> Kuehner, Jason N., and David A. Brow. “Regulation of a Eukaryotic Gene by GTP-Dependent Start Site Selection and Transcription Attenuation.” Molecular Cell, vol. 31, no. 2, 2008, pp. 201–211., doi:10.1016/j.molcel.2008.05.018. </ref> Hrp1 also activates the nonsense-mediated mRNA decay pathway, which monitors translation and degrades incorrect mRNA.<ref> González, Carlos I., et al. “The Yeast HnRNP-like Protein Hrp1/Nab4 Marks a Transcript for Nonsense-Mediated MRNA Decay.” Molecular Cell, vol. 5, no. 3, 2000, pp. 489–499., doi:10.1016/s1097-2765(00)80443-8. </ref> Hrp1, along with Rna14, competes with Npl13, which inhibits cleavage at the polyadenylation site.<ref> Bucheli, M. E., et al. “Polyadenylation Site Choice in Yeast Is Affected by Competition between Npl3 and Polyadenylation Factor CFI.” RNA, vol. 13, no. 10, 2007, pp. 1756–1764., doi:10.1261/rna.607207. </ref> | ||
| - | <StructureSection load='2cjk' size='400' side='right' caption='(PDB entry [[2cjk]])' scene=''> | ||
== Structure == | == Structure == | ||
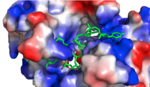
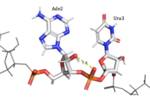
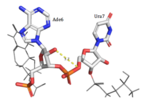
HRP1 is made up of two [https://en.wikipedia.org/wiki/RNA RNA] binding domains (RBDs) that contain residues serving to facilitate RNA recognition. These two domains fold into a βαββαβ [https://en.wikipedia.org/wiki/Protein_secondary_structure secondary structure]<ref>Clery, Antoine, et al. “RNA Recognition Motifs: Boring? Not Quite.” Current Opinion in Structural Biology, Elsevier Current Trends, 30 May 2008, www.sciencedirect.com/science/article/pii/S0959440X08000584.</ref> in an RNA-free environment, allowing Hrp1 to behave rigidly. The <scene name='78/782604/First_rbd/4'>first RBD</scene> includes residues extending from Ser158 to Ala233 and the <scene name='78/782604/Second_rbd/2'>second RBD</scene> extends from Lys244 to Ala318. Both RBDs are composed of a four-stranded [https://en.wikipedia.org/wiki/Beta_sheet beta sheet] with two [https://en.wikipedia.org/wiki/Alpha_helix alpha helices] spanning across one side of the sheet. The linker region is made up of residues spanning from Ile234 to Gly243. When RNA is introduced into the environment, conformational change is demonstrated within the linker region and a <scene name='78/782604/Linker_helix/2'>short two-turn alpha helix</scene> forms from Arg236 to Lys241. The helix that forms is made up of many charged polar residues that [https://en.wikipedia.org/wiki/Salt_bridge_(protein_and_supramolecular) stablilize] themselves through <scene name='78/782604/Salt_bridges/2'>salt bridge interactions</scene> between Arg236-Asp240 and Asp237-Lys241.<ref>Perez-Canadillas, Jose Manuel. “Grabbing the Message: Structural Basis of MRNA 3â²UTR Recognition by Hrp1.” The EMBO Journal, vol. 25, no. 13, 2006, pp. 3167–3178., doi:10.1038/sj.emboj.7601190. <ref name= "rna"> </ref> | HRP1 is made up of two [https://en.wikipedia.org/wiki/RNA RNA] binding domains (RBDs) that contain residues serving to facilitate RNA recognition. These two domains fold into a βαββαβ [https://en.wikipedia.org/wiki/Protein_secondary_structure secondary structure]<ref>Clery, Antoine, et al. “RNA Recognition Motifs: Boring? Not Quite.” Current Opinion in Structural Biology, Elsevier Current Trends, 30 May 2008, www.sciencedirect.com/science/article/pii/S0959440X08000584.</ref> in an RNA-free environment, allowing Hrp1 to behave rigidly. The <scene name='78/782604/First_rbd/4'>first RBD</scene> includes residues extending from Ser158 to Ala233 and the <scene name='78/782604/Second_rbd/2'>second RBD</scene> extends from Lys244 to Ala318. Both RBDs are composed of a four-stranded [https://en.wikipedia.org/wiki/Beta_sheet beta sheet] with two [https://en.wikipedia.org/wiki/Alpha_helix alpha helices] spanning across one side of the sheet. The linker region is made up of residues spanning from Ile234 to Gly243. When RNA is introduced into the environment, conformational change is demonstrated within the linker region and a <scene name='78/782604/Linker_helix/2'>short two-turn alpha helix</scene> forms from Arg236 to Lys241. The helix that forms is made up of many charged polar residues that [https://en.wikipedia.org/wiki/Salt_bridge_(protein_and_supramolecular) stablilize] themselves through <scene name='78/782604/Salt_bridges/2'>salt bridge interactions</scene> between Arg236-Asp240 and Asp237-Lys241.<ref>Perez-Canadillas, Jose Manuel. “Grabbing the Message: Structural Basis of MRNA 3â²UTR Recognition by Hrp1.” The EMBO Journal, vol. 25, no. 13, 2006, pp. 3167–3178., doi:10.1038/sj.emboj.7601190. <ref name= "rna"> </ref> | ||
Revision as of 03:20, 3 April 2018
HRP1 found in Saccharomyces cerevisiae
Introduction
Hrp1 is a heterogeneous ribonuclear protein of Saccharomyces cerevisiae, baker’s yeast. Hrp1 is an essential component of 3’ pre-mRNA processing and contributes to the preparatory cleavage required for polyadenylation. The gene expressed as Hrp1, HRP1, was first isolated by Henry, et al.[1] and was later attributed to the Hrp1 protein by Kessler, et al.[2] Hrp1 also participates in the regulation of the 3’ end.
| |||||||||||