This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Isabella Gieck/Sandbox 1
From Proteopedia
| Line 34: | Line 34: | ||
=== Adenosine recognition === | === Adenosine recognition === | ||
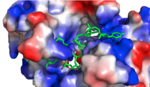
| - | + | Arg321 and a hydrogen bond between N1 and the amide backbone of Ile234. Upon binding, <scene name='78/782604/Adenosine_4_interactions/3'>Adenosine 4</scene> is fit inside of a deep hydrophobic pocket made up of Trp168 and Lys226. The stacking of Trp in this interaction is demonstrated as a unique feature of Hrp1; similar Hrp1-like proteins maintain this conserved Trp, but do not demonstrate Trp stacking. Trp168 also forms a hydrogen bond with N7 of this base. Additionally, Ade4 makes base specific contacts with Lys226 through a pi-cation interaction and with Asn167 as N6 of the base donates a hydrogen. The hydrophobic pocket in which <scene name='78/782604/Adenosine_6_interactions/2'>Adenosine 6 interactions</scene> resides upon binding is made up of Phe162 and Ile234, which sandwich Ade6. N1 of this base forms a hydrogen bond with the amide backbone of Glu319, and N6 acts as a hydrogen donor for Arg232. | |
| - | <scene name='78/782604/Adenosine_6_interactions/2'>Adenosine 6 interactions</scene> | ||
=== Uracil recognition === | === Uracil recognition === | ||
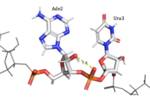
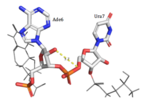
| - | Hrp1 interacts with the three uracil bases mainly though Van der Waals contacts. All three uridines interact with aromatic residues of the protein, although Ura3 and Ura5 have low surface accessibility compared to Ura7. | + | Hrp1 interacts with the three uracil bases mainly though Van der Waals contacts. All three uridines interact with aromatic residues of the protein, although Ura3 and Ura5 have low surface accessibility compared to Ura7. <scene name='78/782604/Uracil_3_interactions/2'>Uracil 3 </scene> interacts with Phe288 in a nonplanar position, and RNA discrimination here is suggested to be mediated by recognition of the imino N3 by the backbone phosphate of Ade6. <scene name='78/782604/Uracil_5_interactions/2'>Uracil 5 </scene> has a similar non-planar ring contact with Phe162, although it is suggested to be a stronger interaction than that between Ura3 and Phe288. Base discrimination at this site relies on the O2 and O4 of Ura5 hydrogen bonding with Lys244 and Lys231, respectively. While these two lysines are conserved in most Hrp1-like proteins, there are variations in other organisms that replace Lys244 with an Asn, although the interaction remains conserved. <scene name='78/782604/Uracil_7_interactions/3'>Uracil 7 </scene> does form a planar stacking arrangement with the aromatic ring of Phe202. Base discrimination here relies heavily on the hydrogen bonding between the O4 of the uracil base and the amine group of Lys160. |
| - | + | ||
| - | + | ||
| - | + | ||
| - | <scene name='78/782604/Uracil_7_interactions/3'>Uracil 7 | + | |
</StructureSection> | </StructureSection> | ||
Revision as of 19:37, 27 April 2018
Contents |
Heterogeneous Ribonucleoprotein 1 (HRP1) found in Saccharomyces cerevisiae
Introduction
Hrp1 is a heterogeneous ribonuclear protein of Saccharomyces cerevisiae, baker’s yeast. Hrp1 is an essential component of 3’ pre-mRNA processing and contributes to the preparatory cleavage required for polyadenylation. The gene expressed as Hrp1, HRP1, was first isolated by Henry, et al.[1] and was later attributed to the Hrp1 protein by Kessler, et al.[2] Hrp1 also participates in the regulation of the 3’ end.
Function
| |||||||||||
Novelty
While Trp168 is a highly conserved residue amongst fungal HRP1-like proteins, the tryptophan stacking found with Ade4 appears to be a novel feature not found in other single-stranded RNA and 2 RBD complexes. It is believed that Trp168 and the ring stacking it engages in are crucial to the affinity and base specificity for RNA.
To research Trp168's importance, studies have experimentally replaced it with phenylalanine and alanine side chains. While these mutants retained the ability to form 1 protein:1 RNA ratio complexes, the affinity for the AU repetition PEE is at least 10 times weaker than that of the wild-type. Furthermore, results of 15N-HSQC spectra comparisons suggest that while the Phe and Ala mutants share similar RNA-binding modes to each, they are completely different from the wild-type Trp.
Another structural finding of the splicing-factor Fox-1 in complex with RNA identifies Phe126 to have an equivalent position as Trp128 has in Hrp1. Studies were done to test the importance of Phe126 in RNA binding by mutating this residue. Similarly to the experiment mentioned above, it was concluded that the aromatic structure of Phe126 played an important role with affinity, as the residue engages in 2 planar stacking interactions with 2 RNA bases and makes contact with a third base. However, aromatic mutants did not have a significant effect on affinity, which suggests that they share a similar binding mode to the Phe126 wild-type. This is not the case with Trp168 in Hrp1, indicating that perhaps Hrp1 has strict sequence requirements.
Disease
Though Hrp1 is not analogous to any mammalian hnRNP[23], the protein and its corresponding gene are occasionally studied as orthologues to human hnRNPs. HNRPDL is one such family of human hnRNPs. Mutations to several members of this class of hnRNPs result in many facets of muscular dystrophy. A study by Vieira, et al. [24] found that elimination of Hrp1 had profound effects on protein localization and activation, and these results were used as a model for the genotypic causation of muscular dystrophy.
References
- ↑ Henry, Michael, et al. “Potential RNA Binding Proteins in Saccharomyces Cerevisiae Identified as Suppressors of Temperature-Sensitive Mutations inNPL3.” Genetics, vol. 142, Jan. 1996, pp. 103–115.
- ↑ Kessler, Marco M, et al. “Purification of the Saccharomyces Cerevisiae Cleavage/Polyadenylation Factor I.” Journal of Biological Chemistry, vol. 271, no. 43, 25 Oct. 1996, pp. 27167–27175.
- ↑ Guisbert, K. Kim. “Functional Specificity of Shuttling HnRNPs Revealed by Genome-Wide Analysis of Their RNA Binding Profiles.” RNA, vol. 11, no. 4, Jan. 2005, pp. 383–393., doi:10.1261/rna.7234205.
- ↑ Kessler, Marco M, et al. “Purification of the Saccharomyces Cerevisiae Cleavage/Polyadenylation Factor I.” Journal of Biological Chemistry, vol. 271, no. 43, 25 Oct. 1996, pp. 27167–27175.
- ↑ Minvielle-Sebastia, L. “Control of Cleavage Site Selection during MRNA 3 End Formation by a Yeast HnRNP.” The EMBO Journal, vol. 17, no. 24, 1998, pp. 7454–7468., doi:10.1093/emboj/17.24.7454.
- ↑ Kessler, M. M., et al. “Hrp1, a Sequence-Specific RNA-Binding Protein That Shuttles between the Nucleus and the Cytoplasm, Is Required for MRNA 3-End Formation in Yeast.” Genes & Development, vol. 11, no. 19, Jan. 1997, pp. 2545–2556., doi:10.1101/gad.11.19.2545.
- ↑ Barnwal, R. P., et al. “Structural and Biochemical Analysis of the Assembly and Function of the Yeast Pre-MRNA 3 End Processing Complex CF I.” Proceedings of the National Academy of Sciences, vol. 109, no. 52, Oct. 2012, pp. 21342–21347., doi:10.1073/pnas.1214102110.
- ↑ Barnwal, R. P., et al. “Structural and Biochemical Analysis of the Assembly and Function of the Yeast Pre-MRNA 3 End Processing Complex CF I.” Proceedings of the National Academy of Sciences, vol. 109, no. 52, Oct. 2012, pp. 21342–21347., doi:10.1073/pnas.1214102110.
- ↑ Leeper, Thomas C., et al. “Novel Protein–Protein Contacts Facilitate MRNA 3′-Processing Signal Recognition by Rna15 and Hrp1.” Journal of Molecular Biology, vol. 401, no. 3, 2010, pp. 334–349., doi:10.1016/j.jmb.2010.06.032.
- ↑ Guisbert, K. Kim. “Functional Specificity of Shuttling HnRNPs Revealed by Genome-Wide Analysis of Their RNA Binding Profiles.” RNA, vol. 11, no. 4, Jan. 2005, pp. 383–393., doi:10.1261/rna.7234205.
- ↑ Chen, S. “A Specific RNA-Protein Interaction at Yeast Polyadenylation Efficiency Elements.” Nucleic Acids Research, vol. 26, no. 21, Jan. 1998, pp. 4965–4974., doi:10.1093/nar/26.21.4965.
- ↑ Minvielle-Sebastia, L. “Control of Cleavage Site Selection during MRNA 3 End Formation by a Yeast HnRNP.” The EMBO Journal, vol. 17, no. 24, 1998, pp. 7454–7468., doi:10.1093/emboj/17.24.7454.
- ↑ Henry, M. F. “The Yeast HnRNP-like Protein Hrp1/Nab4 Accumulates in the Cytoplasm after Hyperosmotic Stress: A Novel Fps1-Dependent Response.” Molecular Biology of the Cell, vol. 14, no. 9, Nov. 2003, pp. 3929–3941., doi:10.1091/mbc.e03-01-0854.
- ↑ Henry, M. F. “The Yeast HnRNP-like Protein Hrp1/Nab4 Accumulates in the Cytoplasm after Hyperosmotic Stress: A Novel Fps1-Dependent Response.” Molecular Biology of the Cell, vol. 14, no. 9, Nov. 2003, pp. 3929–3941., doi:10.1091/mbc.e03-01-0854.
- ↑ Kessler, M. M., et al. “Hrp1, a Sequence-Specific RNA-Binding Protein That Shuttles between the Nucleus and the Cytoplasm, Is Required for MRNA 3-End Formation in Yeast.” Genes & Development, vol. 11, no. 19, Jan. 1997, pp. 2545–2556., doi:10.1101/gad.11.19.2545.
- ↑ Lange, Allison, et al. “A PY-NLS Nuclear Targeting Signal Is Required for Nuclear Localization and Function of TheSaccharomyces CerevisiaemRNA-Binding Protein Hrp1.” Journal of Biological Chemistry, vol. 283, no. 19, 2008, pp. 12926–12934., doi:10.1074/jbc.m800898200.
- ↑ Tuck, Alex C, and David Tollervey. “A Transcriptome-Wide Atlas of RNP Composition Reveals Diverse Classes of MRNAs and LncRNAs.” Cell, vol. 154, 29 Aug. 2013, pp. 996–1009.
- ↑ Kuehner, Jason N., and David A. Brow. “Regulation of a Eukaryotic Gene by GTP-Dependent Start Site Selection and Transcription Attenuation.” Molecular Cell, vol. 31, no. 2, 2008, pp. 201–211., doi:10.1016/j.molcel.2008.05.018.
- ↑ González, Carlos I., et al. “The Yeast HnRNP-like Protein Hrp1/Nab4 Marks a Transcript for Nonsense-Mediated MRNA Decay.” Molecular Cell, vol. 5, no. 3, 2000, pp. 489–499., doi:10.1016/s1097-2765(00)80443-8.
- ↑ Bucheli, M. E., et al. “Polyadenylation Site Choice in Yeast Is Affected by Competition between Npl3 and Polyadenylation Factor CFI.” RNA, vol. 13, no. 10, 2007, pp. 1756–1764., doi:10.1261/rna.607207.
- ↑ Clery, Antoine, et al. “RNA Recognition Motifs: Boring? Not Quite.” Current Opinion in Structural Biology, Elsevier Current Trends, 30 May 2008, www.sciencedirect.com/science/article/pii/S0959440X08000584.
- ↑ Perez-Canadillas, Jose Manuel. “Grabbing the Message: Structural Basis of MRNA 3â²UTR Recognition by Hrp1.” The EMBO Journal, vol. 25, no. 13, 2006, pp. 3167–3178., doi:10.1038/sj.emboj.7601190.
- ↑ Gross, S., and C. Moore. “Five Subunits Are Required for Reconstitution of the Cleavage and Polyadenylation Activities of Saccharomyces Cerevisiae Cleavage Factor I.” Proceedings of the National Academy of Sciences, vol. 98, no. 11, Aug. 2001, pp. 6080–6085., doi:10.1073/pnas.101046598.
- ↑ Vieira, Natássia M., et al. “A Defect in the RNA-Processing Protein HNRPDL Causes Limb-Girdle Muscular Dystrophy 1G (LGMD1G).” Human Molecular Genetics, vol. 23, no. 15, 2014, pp. 4103–4110., doi:10.1093/hmg/ddu127.