Here are two ways of visualizing the position of a lipid bilayer membrane around the integral membrane portion of a trans-membrane protein.
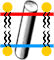 |
Outside
Inside
|
- 1. represent the water-lipid boundaries. Red pseudoatoms represent the extracellular boundary, and blue pseudoatoms represent the cytoplasmic boundary.[1]
- 2. A represents membrane near the protein. If desired, .
a hydrophobic vs. polar color scheme, the protein within the lipid bilayer is seen to have a largely hydrophobic surface. The example used here is 5lil.
Methods: Pseudoatoms
Any protein structure that includes an integral membrane domain can have pseudoatoms added to the PDB file by the Orientations of Proteins in Membranes Server[1]. Structures in the Protein Data Bank are pre-processed; other structures may be uploaded to the server. The server removes hydrogen atoms from the model.
You can download the pseudoatom-enhanced PDB file from the server, and then upload it to Proteopedia for use in a Molecular Scene.
In the case of 5LiL, I had difficulty getting the pseudoatoms to show from a green link (even though they displayed in the SAT). In the PDB file, the pseudoatoms followed a MASTER record. I deleted the MASTER record and all CONECT records, and then I got the results shown above with Image:5lil opm2.pdb.
The pseudoatoms have residue name DUM (for dummy atoms). This can be used in the SAT to select them. The red pseudoatoms are oxygens, and the blue pseudoatoms are nitrogens. Thus, coloring them by chemical element (CPK) will make them red and blue.
Methods: Translucent Cylinder
Choose boundary residues
First you must choose residues or atoms in the model that are closest to the ends of the desired cylinder. If you wish, you can use a pseudoatom-enhanced model to assist.
Viewing the model in FirstGlance in Jmol, you can use Hydrophobic/Polar (Views tab) to visualize the domain with a hydrophobic surface. Touching or clicking on atoms at the domain boundaries will identify those amino acids. If you wish, you can center a particular residue, then zoom in, and see atomic detail with Vines/Sticks (Views tab).
For 5LiL, I chose thr607 and leu250. These will be used in the example commands below, but you must substitute appropriate amino acids for your model.
5LiL is a homo-dimer of chains A and B. Since I did not specify a chain in the commands below, the position of, for example, thr607 will be the average position of all the atoms in thr607 in both chains. That average position will be at the axis of the rod-shaped molecule. Your molecule may require a different strategy.
Generate the Cylinder
To generate the cylinder in the SAT, click on the button below JSmol Advanced: Open JSmol Console. The command to generate the cylinder will have this form:
draw cyl1 cylinder diameter 100.0 color gray (thr607) (leu250)
Adapt this command to your situation, copy and paste it into the LOWER box of the Console, and click the Run button at the lower-left of the console box.
- cyl1 is an arbitrary name for this cylinder.
- You may adjust the diameter and color to your taste. (Gray is a standard color for hydrophobic. Yellow is sometimes used with the mnemonic butter in mind.)
- If you wish to delete all cylinders before making a new one, the command is
draw * delete
- To make the cylinder translucent, the command is
draw cyl1 color translucent -1
Once you have the desired cylinder in the SAT, save the scene, insert a green link in the wikitext box, and save the page. Then test the green link.
After displaying the cylinder with your new green link, if you then click a different green link, the cylinder may persist, unwanted. To fix this:
- In the SAT, load the other scene. It doesn't matter if the cylinder is not visible in the SAT.
- Open the command console.
- Enter and Run the above delete command.
- Save the scene.
- Update the green link to use the newly saved version of the scene.
- Test.
Color The Cylinder Ends
Here are the commands to generate a translucent cylinder with a red top and a blue bottom:
draw cyl1 cylinder diameter 100.0 color gray (thr607) (leu250) nofill mesh
draw cyl1 color translucent -1
draw ctop circle diameter 100.0 color red (thr607) (leu250)
draw ctop color translucent -1
draw cbot circle diameter 100.0 color blue (leu250) (thr607)
draw cbot color translucent -1
- Adding nofill mesh to the end of the first draw command removes the ends of the gray cylinder.
- Next, the two circle commands create new cylinder ends (circles) colored red and blue.
- Notice the reversed order of the amino acids in the two circle commands. The first position is the center of the circle. The second position makes the circle perpindicular to the axis between the two specified positions.
If you have questions or need more details, don't hesitate to email 

