This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
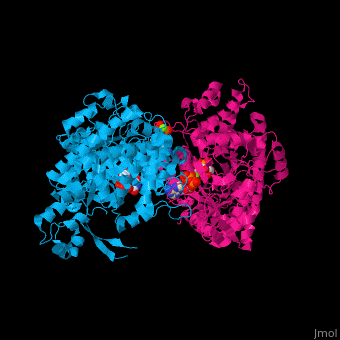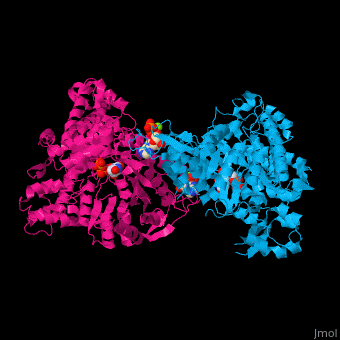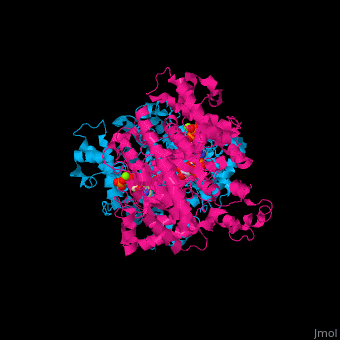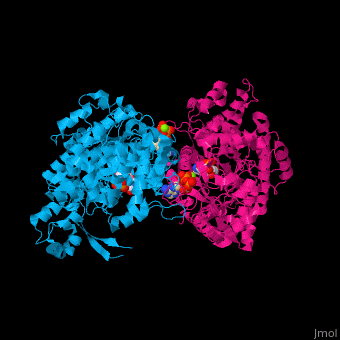
Ribonucleotide reductase
From Proteopedia
(Difference between revisions)
| Line 2: | Line 2: | ||
== Function == | == Function == | ||
'''Ribonucleotide reductase''' (RNR) or '''ribonucleotide-diphosphate reductase''' catalyzes the formation of deoxyribonucleotides from ribonucleotides<ref>PMID:16756507</ref>. There are 3 classes of RNR.<br /> | '''Ribonucleotide reductase''' (RNR) or '''ribonucleotide-diphosphate reductase''' catalyzes the formation of deoxyribonucleotides from ribonucleotides<ref>PMID:16756507</ref>. There are 3 classes of RNR.<br /> | ||
| - | * '''Class | + | * '''Class Ia RNR''' is a tetramer composed from large (RNR1) and small (RNR2) subunits. Class I RNR is iron-dependent and produces tyrosyl radical. Thimidine triphosphate (TTP) is an effector in the reaction<ref>PMID:21456967</ref>.<br /> |
* ''' Class Ib RNR''' contains 2 proteins: α (NrdE) and β (NrdF)<ref>PMID:21250660</ref>.<br /> | * ''' Class Ib RNR''' contains 2 proteins: α (NrdE) and β (NrdF)<ref>PMID:21250660</ref>.<br /> | ||
| + | ''E. Coli'' contains two types of RNR: the aerobic RNR belongs to class Ia and is composed of proteins R1 and R2, the anaerobic RNR belongs to class Ib. <br /> | ||
* ''' Class II RNR''' reduces ribonucleotide triphosphates using coenzyme B12<ref>PMID:11875520</ref>.<br /> | * ''' Class II RNR''' reduces ribonucleotide triphosphates using coenzyme B12<ref>PMID:11875520</ref>.<br /> | ||
* '''Class III RNR''' generate glycine radical using S-adenosyl methionine and Fe-S center<ref>PMID:17606358</ref>.<br /> | * '''Class III RNR''' generate glycine radical using S-adenosyl methionine and Fe-S center<ref>PMID:17606358</ref>.<br /> | ||
| Line 29: | Line 30: | ||
**[[1zyz]], [[1zzd]], [[2cvs]], [[2cvt]], [[2cvu]], [[2cvv]], [[2cvw]], [[2cvx]], [[2cvy]] - yRNR R1 – yeast<BR /> | **[[1zyz]], [[1zzd]], [[2cvs]], [[2cvt]], [[2cvu]], [[2cvv]], [[2cvw]], [[2cvx]], [[2cvy]] - yRNR R1 – yeast<BR /> | ||
| - | **[[1r1r | + | **[[1r1r]] - EcRNR R1 – ''Escherichia coli''<br /> |
| - | **[[5r1r]], [[6r1r]], [[7r1r]] | + | **[[1mrr]] - EcRNR R1 + Mn<br /> |
| - | + | **[[5r1r]], [[6r1r]], [[7r1r]], [[1r1r]] – EcRNR R1 (mutant) + R2<br /> | |
*''Ribonucleotide reductase large subunit complex'' | *''Ribonucleotide reductase large subunit complex'' | ||
| Line 50: | Line 51: | ||
**[[2zlf]], [[2zlg]] - yRNR R1 + peptide<BR /> | **[[2zlf]], [[2zlg]] - yRNR R1 + peptide<BR /> | ||
**[[2eud]] - yRNR R1 + ligand<BR /> | **[[2eud]] - yRNR R1 + ligand<BR /> | ||
| - | **[[3rsr]] – yRNR + inhibitor<br | + | **[[3rsr]] – yRNR R1 + inhibitor<br /> |
| - | + | ||
| - | + | ||
| - | + | ||
**[[1qfn]] - EcRNR R1 + glutaredoxin 1<br /> | **[[1qfn]] - EcRNR R1 + glutaredoxin 1<br /> | ||
| - | **[[1peu]] - StRNR + DTP - ''Salmonella typhimurium''<br /> | ||
*RNR small subunit | *RNR small subunit | ||
| - | **[[2xof]], [[ | + | **[[2xof]], [[1av8]], [[1rib]] – EcRNR R2 + Fe2O<BR /> |
| - | ** | + | **[[2alx]], [[1jqc]], [[1jpr]] - EcRNR R2 + Mn<br /> |
| + | **[[1piy]], [[1r65]], [[1mxr]], [[1xik]] - EcRNR R2 + Fe<br /> | ||
| + | **[[1piz]], [[1pj0]], [[1pj1]], [[1pim]], [[1piu]], [[1biq]], [[1pfr]] - EcRNR R2 (mutant) + Fe<BR /> | ||
| + | **[[1yfd]], [[2av8]] - EcRNR R2 (mutant) + Fe2O<br /> | ||
| + | **[[1pm2]] - EcRNR R2 (mutant) + Mn<br /> | ||
**[[3hf1]], [[2vux]], [[2uw2]], [[4djn]] – hRNR M2<BR /> | **[[3hf1]], [[2vux]], [[2uw2]], [[4djn]] – hRNR M2<BR /> | ||
| - | **[[3mjo]], [[3dhz]], [[1oqu]], [[1kgn]], [[1kgo]], [[1kgp]] – RNR R2 – ''Corynebacterium ammoniagenes''<BR /> | ||
**[[2p1i]] – RNR R2 – ''Plasmodium yoelii''<BR /> | **[[2p1i]] – RNR R2 – ''Plasmodium yoelii''<BR /> | ||
**[[2o1z]] - RNR R2 – ''Plasmodium vivax''<BR /> | **[[2o1z]] - RNR R2 – ''Plasmodium vivax''<BR /> | ||
| Line 85: | Line 85: | ||
*''Ribonucleotide reductase small subunit complex'' | *''Ribonucleotide reductase small subunit complex'' | ||
| - | **[[1rsr]], [[1rsv]] - EcRNR R2 (mutant) + N3 | + | **[[1rsr]], [[1rsv]] - EcRNR R2 (mutant) + Fe + N3<BR /> |
| - | + | **[[1rnr]] – EcRNR R2 (mutant) + Fe + dopa<BR /> | |
| - | **[[1rnr]] – EcRNR R2 (mutant) + dopa<BR /> | + | **[[5ci4]] – EcRNR R2 (mutant) + Fe2O<br /> |
| - | **[[5ci4]] – EcRNR R2 (mutant) + | + | **[[5ci3]], [[5ci0]], [[5ci1]], [[5ci2]] – EcRNR R2 (mutant) + fluoro-tyrosine + Fe<sub>2</sub>O<br /> |
| - | **[[5ci3]], [[5ci0]], [[5ci1]], [[5ci2]] – EcRNR R2 (mutant) + fluoro-tyrosine + | + | |
**[[1w69]] – mRNR R2 + acetate<br /> | **[[1w69]] – mRNR R2 + acetate<br /> | ||
**[[6cwp]] – FjRNR + Mn + peptide<br /> | **[[6cwp]] – FjRNR + Mn + peptide<br /> | ||
| Line 95: | Line 94: | ||
*Ribonucleotide reductase small+large subunit | *Ribonucleotide reductase small+large subunit | ||
| - | **[[3uus]] | + | **[[3uus]] - EcRNR R1+ R2 + Fe + ATP<br /> |
| - | **[[4erp]] - EcRNR R1+ R2 + | + | **[[4erm]] - EcRNR R1+ R2 + Fe2O + ATP + ADP<br /> |
| - | **[[2xak]], [[2xap]], [[2xo4]], [[2xo5]] - EcRNR R1+ R2 peptide<BR /> | + | **[[4erp]] - EcRNR R1+ R2 + Fe2O + ATP<br /> |
| - | **[[2xav]], [[2xaw]], [[2xax]], [[2xay]], [[2xaz]], [[2x0x]] - EcRNR R1 (mutant) + R2 peptide<BR /> | + | **[[2xak]], [[2xap]], [[2xo4]], [[2xo5]] - EcRNR R1 modified+ R2 peptide<BR /> |
| - | **[[5cns]], [[5cnt]], [[5cnv]], [[5cnu]] - EcRNR R1 + R2 + 2 nucleotides<br /> | + | **[[2xav]], [[2xaw]], [[2xax]], [[2xay]], [[2xaz]], [[2x0x]] - EcRNR R1 (mutant) modified + R2 peptide<BR /> |
| + | **[[5cns]], [[5cnt]], [[5cnv]], [[5cnu]] - EcRNR R1 + R2 + Fe2O + 2 nucleotides<br /> | ||
| + | **[[2r1r]] - EcRNR R1 + R2 + TTP<BR /> | ||
| + | **[[3r1r]] - EcRNR R1 + R2 + AMPPNP<BR /> | ||
| + | **[[4r1r]] - EcRNR R1 + R2 + GDP + TTP<BR /> | ||
**[[2bq1]] - StRNR R1+ R2<br /> | **[[2bq1]] - StRNR R1+ R2<br /> | ||
**[[5im3]] - RNR + dATP – ''Pseudomonas aeruginosa''<br /> | **[[5im3]] - RNR + dATP – ''Pseudomonas aeruginosa''<br /> | ||
| Line 107: | Line 110: | ||
**[[6cgm]] - BaRNR subunit NrdE – ''Bacillus subtilis''<br /> | **[[6cgm]] - BaRNR subunit NrdE – ''Bacillus subtilis''<br /> | ||
**[[6cgn]], [[6cgl]] - BaRNR subunit NrdE + AMP derivative <br /> | **[[6cgn]], [[6cgl]] - BaRNR subunit NrdE + AMP derivative <br /> | ||
| + | **[[1pem]] - StRNR NrdE – Salmonella typhimurium<br /> | ||
| + | **[[1peo]], [[1peq]], [[1peu]] - StRNR NrdE + nucleotide<br /> | ||
| + | **[[2bq1]] - StRNR NrdE + NrdF + Fe + GTP<br /> | ||
**[[4dr0]] - BaRNR subunit NrdF + Mn <br /> | **[[4dr0]] - BaRNR subunit NrdF + Mn <br /> | ||
**[[4bmq]], [[4bmr]], [[4bmt]] - BcRNR NrdF + Fe – ''Bacillus cereus''<br /> | **[[4bmq]], [[4bmr]], [[4bmt]] - BcRNR NrdF + Fe – ''Bacillus cereus''<br /> | ||
**[[4bmu]] - BcRNR NrdF + Mn<br /> | **[[4bmu]] - BcRNR NrdF + Mn<br /> | ||
**[[4bmo]], [[4bmp]] - BcRNR NrdF + NrdI + Fe + FMN<br /> | **[[4bmo]], [[4bmp]] - BcRNR NrdF + NrdI + Fe + FMN<br /> | ||
| + | **[[4m1f]] - EcRNR NrdF<br /> | ||
| + | **[[3n37]] - EcRNR NrdF + Mn<br /> | ||
| + | **[[3n38]] - EcRNR NrdF + Fe<br /> | ||
| + | **[[3n39]], [[3n3a]] - EcRNR NrdF + NrdI + Mn + FMN<br /> | ||
| + | **[[3n3b]] - EcRNR NrdF + NrdI + Mn + FMN + Fe2O<br /> | ||
| + | **[[4n82]] – SsRNR + FMN – ''Streptococcus sanguinis''<br /> | ||
| + | **[[4n83]] - SsRNR NrdF + Mn<br /> | ||
| + | **[[1r2f]] - StRNR NrdF + Fe<br /> | ||
| + | **[[2r2f]] - StRNR NrdF + Fe2O<br /> | ||
| + | **[[3mjo]] - CcRNR NrdF + Mn - ''Corynebacterium ammoniagenes'' <br /> | ||
| + | **[[1kgn]], [[1kgo]], [[1kgp]], [[1oqu]], [[3dhz]] - CcRNR NrdF + Fe<br /> | ||
*'''Class II ribonucleotide reductase''' | *'''Class II ribonucleotide reductase''' | ||
| Line 122: | Line 139: | ||
**[[3o0o]] - TmRNR + TTP + GDP + Mg + deoxyadenosine + B12<br /> | **[[3o0o]] - TmRNR + TTP + GDP + Mg + deoxyadenosine + B12<br /> | ||
**[[3o0q]] - TmRNR + TTP + GDP + Mg + adenosine <br /> | **[[3o0q]] - TmRNR + TTP + GDP + Mg + adenosine <br /> | ||
| - | **[[1pem]] - StRNR R1 <BR /> | ||
| - | **[[1r2f]], [[2r2f]] - StRNR R2<br /> | ||
| - | **[[1peo]], [[1peq]], [[2peu]] - StRNR R1 + effector<BR /> | ||
**[[1l1l]] – RNR (mutant) – ''Lactobacillus leichmannii''<br /> | **[[1l1l]] – RNR (mutant) – ''Lactobacillus leichmannii''<br /> | ||
Revision as of 08:39, 16 October 2018
| |||||||||||
3D Structures of Ribonucleotide reductase
Updated on 16-October-2018
Class Ib RNR
References
- ↑ Nordlund P, Reichard P. Ribonucleotide reductases. Annu Rev Biochem. 2006;75:681-706. PMID:16756507 doi:http://dx.doi.org/10.1146/annurev.biochem.75.103004.142443
- ↑ Cotruvo JA, Stubbe J. Class I ribonucleotide reductases: metallocofactor assembly and repair in vitro and in vivo. Annu Rev Biochem. 2011;80:733-67. doi: 10.1146/annurev-biochem-061408-095817. PMID:21456967 doi:http://dx.doi.org/10.1146/annurev-biochem-061408-095817
- ↑ Cotruvo JA, Stubbe J. Escherichia coli class Ib ribonucleotide reductase contains a dimanganese(III)-tyrosyl radical cofactor in vivo. Biochemistry. 2011 Mar 15;50(10):1672-81. doi: 10.1021/bi101881d. Epub 2011 Feb, 15. PMID:21250660 doi:http://dx.doi.org/10.1021/bi101881d
- ↑ Sintchak MD, Arjara G, Kellogg BA, Stubbe J, Drennan CL. The crystal structure of class II ribonucleotide reductase reveals how an allosterically regulated monomer mimics a dimer. Nat Struct Biol. 2002 Apr;9(4):293-300. PMID:11875520 doi:http://dx.doi.org/10.1038/nsb774
- ↑ Kirdis E, Jonsson IM, Kubica M, Potempa J, Josefsson E, Masalha M, Foster SJ, Tarkowski A. Ribonucleotide reductase class III, an essential enzyme for the anaerobic growth of Staphylococcus aureus, is a virulence determinant in septic arthritis. Microb Pathog. 2007 Nov-Dec;43(5-6):179-88. doi: 10.1016/j.micpath.2007.05.008., Epub 2007 May 25. PMID:17606358 doi:http://dx.doi.org/10.1016/j.micpath.2007.05.008
- ↑ Munro JB, Silva JC. Ribonucleotide reductase as a target to control apicomplexan diseases. Curr Issues Mol Biol. 2012;14(1):9-26. Epub 2011 Jul 26. PMID:21791713
- ↑ Larsson KM, Jordan A, Eliasson R, Reichard P, Logan DT, Nordlund P. Structural mechanism of allosteric substrate specificity regulation in a ribonucleotide reductase. Nat Struct Mol Biol. 2004 Nov;11(11):1142-9. Epub 2004 Oct 10. PMID:15475969 doi:http://dx.doi.org/10.1038/nsmb838
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, Joel L. Sussman, Jaime Prilusky