This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1481
From Proteopedia
| Line 11: | Line 11: | ||
[http://www.rcsb.org/structure/5K7M] | [http://www.rcsb.org/structure/5K7M] | ||
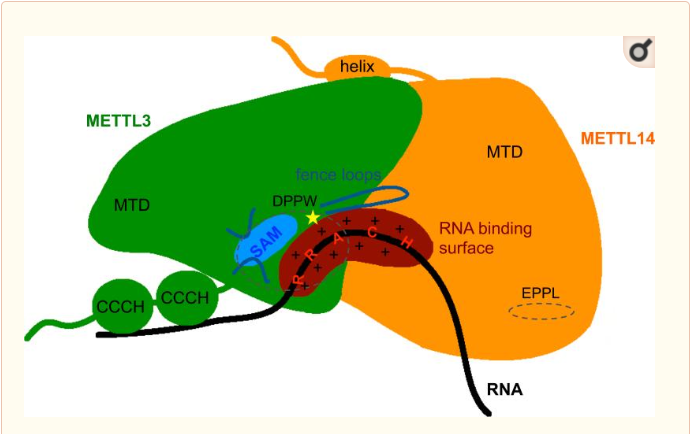
| - | METTL3 and MTTL14 have both a methyltranferase domain but the complex METTL3/METTL14 has a better methyltransferase activity. Moreover, a mutatio in the catalytic center of METTL3 inhibits the hole methyltransferase activity of the complex, whereas a mutation in the catalytic center of METTL14 does not, | + | METTL3 and MTTL14 have both a methyltranferase domain but the complex METTL3/METTL14 has a better methyltransferase activity. Moreover, a mutatio in the catalytic center of METTL3 inhibits the hole methyltransferase activity of the complex, whereas a mutation in the catalytic center of METTL14 does not. Thus, METTL3 is the catalytic subunit of the complex and METTL14 enhances the methyltransfearse activity by stabilizing the complex and enables the recognition of consensus sequence on messenger RNA. |
[[Image:MET.PNG]] | [[Image:MET.PNG]] | ||
| Line 63: | Line 63: | ||
== Complex METTL3/METTL14 enzymatic working process == | == Complex METTL3/METTL14 enzymatic working process == | ||
| + | |||
The enzymatic complex of two proteins is abble to realize methylation reactions thanks to its sequence and specific which allow it to catalyze some perticular reactions at given sites. | The enzymatic complex of two proteins is abble to realize methylation reactions thanks to its sequence and specific which allow it to catalyze some perticular reactions at given sites. | ||
Revision as of 21:05, 28 December 2018
| This Sandbox is Reserved from 06/12/2018, through 30/06/2019 for use in the course "Structural Biology" taught by Bruno Kieffer at the University of Strasbourg, ESBS. This reservation includes Sandbox Reserved 1480 through Sandbox Reserved 1543. |
To get started:
More help: Help:Editing |
Crystal structure of the catalytic domains of Mettl3/Mettl14 complexInsert caption here
Drag the structure with the mouse to rotate
Insert caption here |
| Drag the structure with the mouse to rotate |
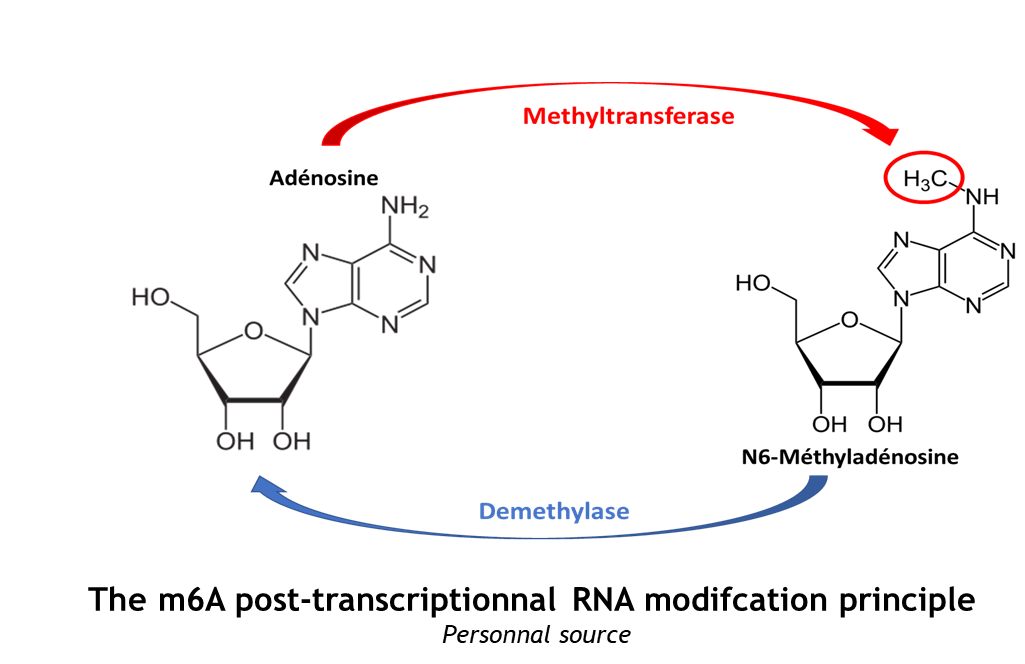
The complex METTL3/METTL14 is a heterodimer enzymatic complex involved into RNA post-transcription modifications by humans. This complex is abble to add a methyl group on adenosin of the RNA, by catalyzing a m6(A) modification.The N(6)-methyladenosine (m(6)A) is a quite common, reversible chemical modifications of RNAs molecules which plays a key role in several biological fonctions. This post transcriptional modification can be added by WRITERS, recognized by READERS and also removed byr ERASERS. The METTL3/METTL14 complex plays the role of writer.
| |||||||||||
References
<Structural Basis for Cooperative Function of Mettl3 and Mettl14 Methyltransferases/>[1] <Structural basis of N(6)-adenosine methylation by the METTL3-METTL14 complex./>[2]
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644