This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1481
From Proteopedia
| Line 4: | Line 4: | ||
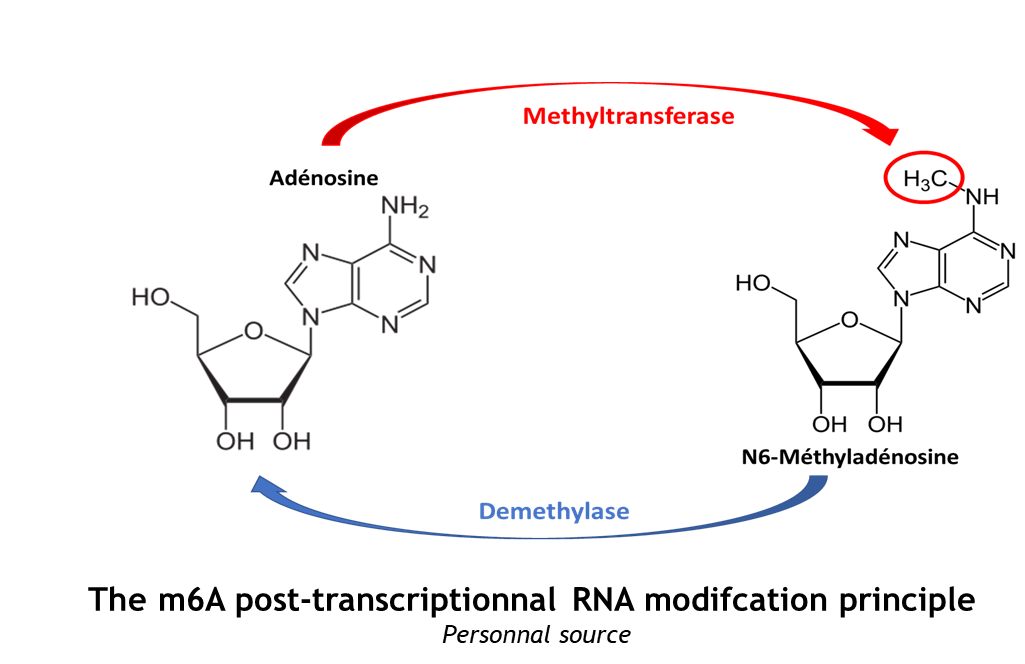
This complex is abble to add a methyl group on adenosin of the RNA, by catalyzing a m6(A) modification.The N(6)-methyladenosine (m(6)A) is a quite common, reversible chemical modification of RNAs molecules, which plays a key role in several different biological fonctions. This post-transcriptional modification can be added by WRITERS, recognized by READERS and also removed byr ERASERS. The METTL3/METTL14 complex plays the role of writer. | This complex is abble to add a methyl group on adenosin of the RNA, by catalyzing a m6(A) modification.The N(6)-methyladenosine (m(6)A) is a quite common, reversible chemical modification of RNAs molecules, which plays a key role in several different biological fonctions. This post-transcriptional modification can be added by WRITERS, recognized by READERS and also removed byr ERASERS. The METTL3/METTL14 complex plays the role of writer. | ||
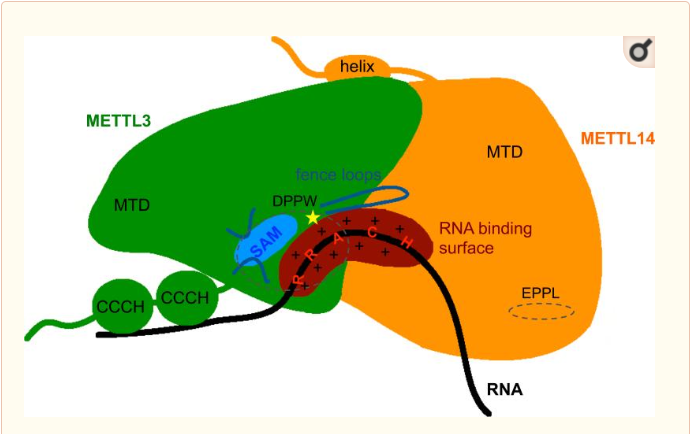
| - | This enzymatic complex belongs to the second class of | + | This enzymatic complex belongs to the second class of enzymes, which are the transferases. The complex is formed by 574 amino acid residues, divided into two different proteins nammed as Methyltransferase Like number 3 and 14. [[Image:writers.png]] |
| - | <StructureSection load=' | + | <StructureSection load='5K7M' size='340' side='right' caption='Caption for this structure' scene=''> |
You may include any references to papers as in: the use of JSmol in Proteopedia <ref>DOI </ref> or to the article describing Jmol <ref>PMID:</ref> to the rescue. | You may include any references to papers as in: the use of JSmol in Proteopedia <ref>DOI </ref> or to the article describing Jmol <ref>PMID:</ref> to the rescue. | ||
Revision as of 16:48, 29 December 2018
| This Sandbox is Reserved from 06/12/2018, through 30/06/2019 for use in the course "Structural Biology" taught by Bruno Kieffer at the University of Strasbourg, ESBS. This reservation includes Sandbox Reserved 1480 through Sandbox Reserved 1543. |
To get started:
More help: Help:Editing |
Crystal structure of the catalytic domains of Mettl3/Mettl14 complexCrystal structure of the catalytic domains of Mettl3/Mettl14 complex
Drag the structure with the mouse to rotate
Crystal structure of the catalytic domains of Mettl3/Mettl14 complex |
| Drag the structure with the mouse to rotate |
The complex METTL3/METTL14 is a heterodimer enzymatic complex involved into RNA post-transcriptional modifications by humans. This complex is abble to add a methyl group on adenosin of the RNA, by catalyzing a m6(A) modification.The N(6)-methyladenosine (m(6)A) is a quite common, reversible chemical modification of RNAs molecules, which plays a key role in several different biological fonctions. This post-transcriptional modification can be added by WRITERS, recognized by READERS and also removed byr ERASERS. The METTL3/METTL14 complex plays the role of writer.
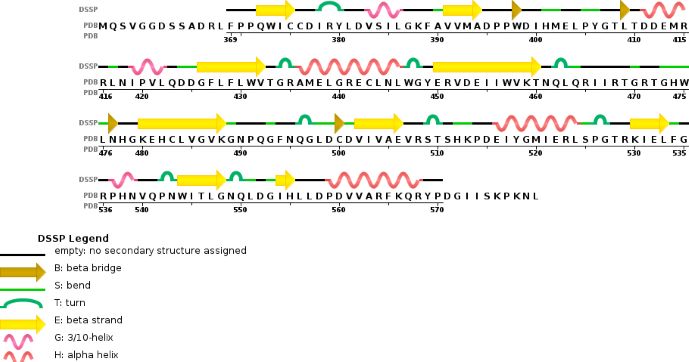
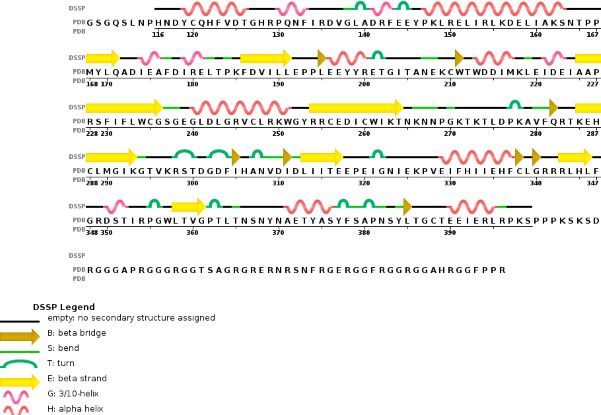
This enzymatic complex belongs to the second class of enzymes, which are the transferases. The complex is formed by 574 amino acid residues, divided into two different proteins nammed as Methyltransferase Like number 3 and 14. 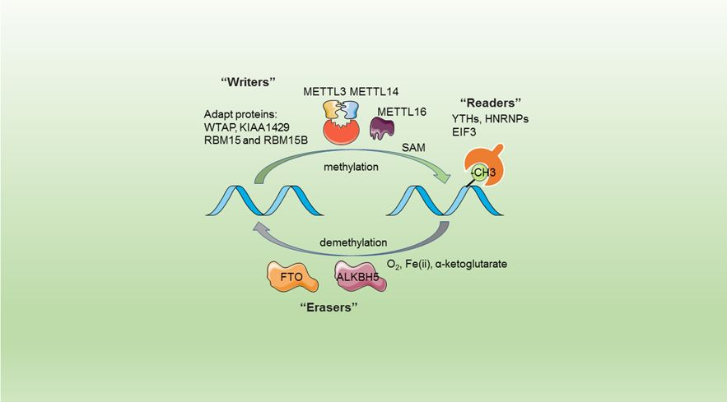
| |||||||||||
References
- ↑ . PMID:216315890657
- ↑ . PMID:216315890657
- ↑ Wang P, Doxtader KA, Nam Y. Structural Basis for Cooperative Function of Mettl3 and Mettl14 Methyltransferases. Mol Cell. 2016 Jul 21;63(2):306-17. doi: 10.1016/j.molcel.2016.05.041. Epub 2016 , Jun 30. PMID:27373337 doi:http://dx.doi.org/10.1016/j.molcel.2016.05.041
- ↑ Wang X, Feng J, Xue Y, Guan Z, Zhang D, Liu Z, Gong Z, Wang Q, Huang J, Tang C, Zou T, Yin P. Structural basis of N(6)-adenosine methylation by the METTL3-METTL14 complex. Nature. 2016 May 25;534(7608):575-8. doi: 10.1038/nature18298. PMID:27281194 doi:http://dx.doi.org/10.1038/nature18298
- ↑ Sledz P, Jinek M. Structural insights into the molecular mechanism of the m(6)A writer complex. Elife. 2016 Sep 14;5. pii: e18434. doi: 10.7554/eLife.18434. PMID:27627798 doi:http://dx.doi.org/10.7554/eLife.18434
- ↑ Wang X, Huang J, Zou T, Yin P. Human m(6)A writers: Two subunits, 2 roles. RNA Biol. 2017 Mar 4;14(3):300-304. doi: 10.1080/15476286.2017.1282025. Epub 2017, Jan 25. PMID:28121234 doi:http://dx.doi.org/10.1080/15476286.2017.1282025
- ↑ doi: https://dx.doi.org/10.2210/pdb5K7M/pdb