This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1489
From Proteopedia
(Difference between revisions)
| Line 17: | Line 17: | ||
== '''Structural highlights''' == | == '''Structural highlights''' == | ||
| - | == Global structure == | + | ===Global structure=== |
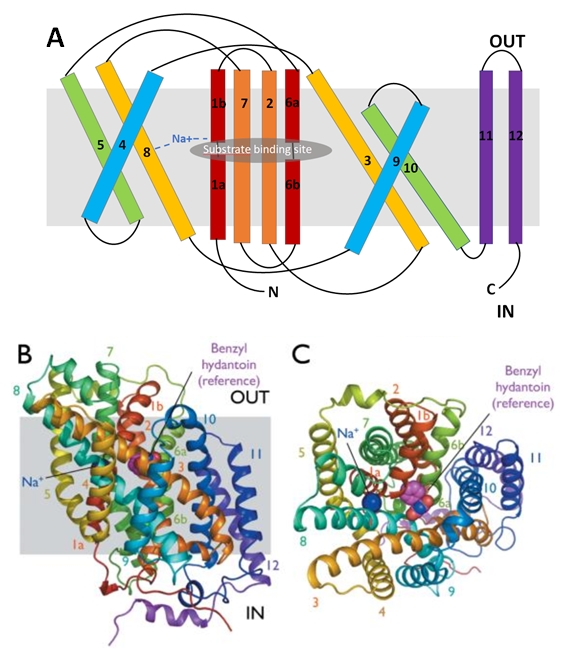
The protein is composed of one chain of 12 transmembrane alpha helices (TMs). They are organised in two repeating units connected by a 59-residue loop (TMs 1-5 and TMs 6-10) and two additional helices (TM 11 and 12). The two repeating units have a symmetrical topology (''Figure 2''). | The protein is composed of one chain of 12 transmembrane alpha helices (TMs). They are organised in two repeating units connected by a 59-residue loop (TMs 1-5 and TMs 6-10) and two additional helices (TM 11 and 12). The two repeating units have a symmetrical topology (''Figure 2''). | ||
| Line 31: | Line 31: | ||
| - | == Structure of the substrate binding site == | + | ===Structure of the substrate binding site=== |
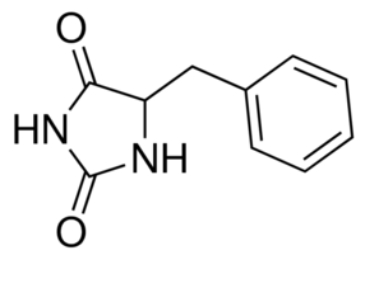
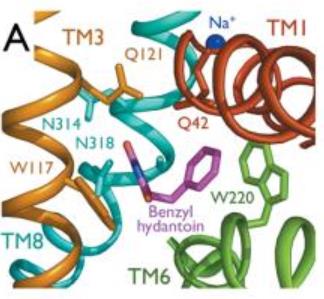
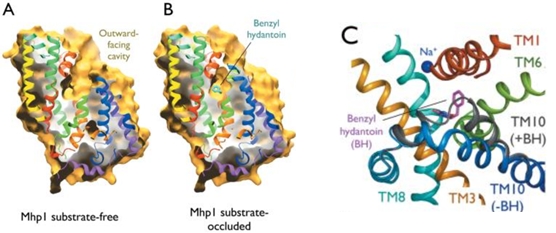
The substrate binding site is located at the break of the TMs 1 and 6. | The substrate binding site is located at the break of the TMs 1 and 6. | ||
| Line 42: | Line 42: | ||
| - | == Structure of the cation binding site == | + | ===Structure of the cation binding site=== |
Mhp1 is a sodium dependent protein. The sodium binds at the C-terminal end of TM1a and interacts with TM8 (''Figure 2.A''). The dipole moment at the C-terminus of TM1a contributes to the binding. | Mhp1 is a sodium dependent protein. The sodium binds at the C-terminal end of TM1a and interacts with TM8 (''Figure 2.A''). The dipole moment at the C-terminus of TM1a contributes to the binding. | ||
Revision as of 18:18, 9 January 2019
| This Sandbox is Reserved from 06/12/2018, through 30/06/2019 for use in the course "Structural Biology" taught by Bruno Kieffer at the University of Strasbourg, ESBS. This reservation includes Sandbox Reserved 1480 through Sandbox Reserved 1543. |
To get started:
More help: Help:Editing |
2JLN
| |||||||||||
References
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644