This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1502
From Proteopedia
(Difference between revisions)
| Line 33: | Line 33: | ||
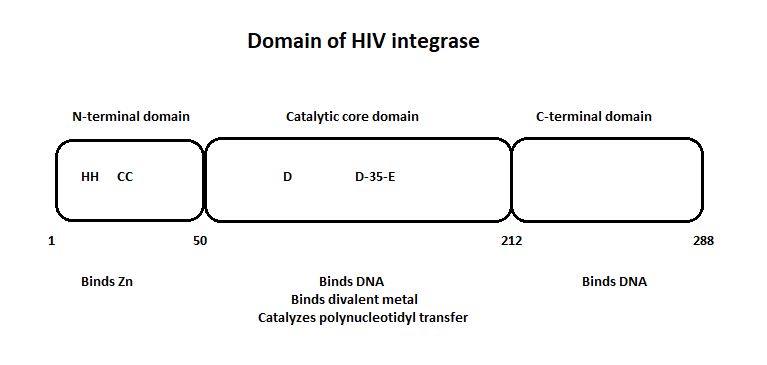
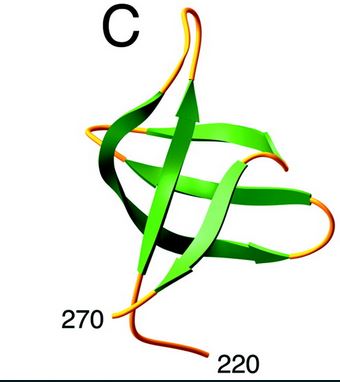
HIV-1 integrase is composed of three domains: the N-terminal (residues 1-49), the core domain (residues 50-212) and the C-terminal domain (residues 213-288). The core domain is responsible for the catalytic activity of the enzyme. It contains three acidic residues, the D,D-35-E motif which plays a key role in catalysis. The N-terminal domain includes the conserved HHCC motif, which binds zinc. The C-terminal domain is less well conserved. [http://www.jbc.org/content/276/26/23213/F2.expansion.html] | HIV-1 integrase is composed of three domains: the N-terminal (residues 1-49), the core domain (residues 50-212) and the C-terminal domain (residues 213-288). The core domain is responsible for the catalytic activity of the enzyme. It contains three acidic residues, the D,D-35-E motif which plays a key role in catalysis. The N-terminal domain includes the conserved HHCC motif, which binds zinc. The C-terminal domain is less well conserved. [http://www.jbc.org/content/276/26/23213/F2.expansion.html] | ||
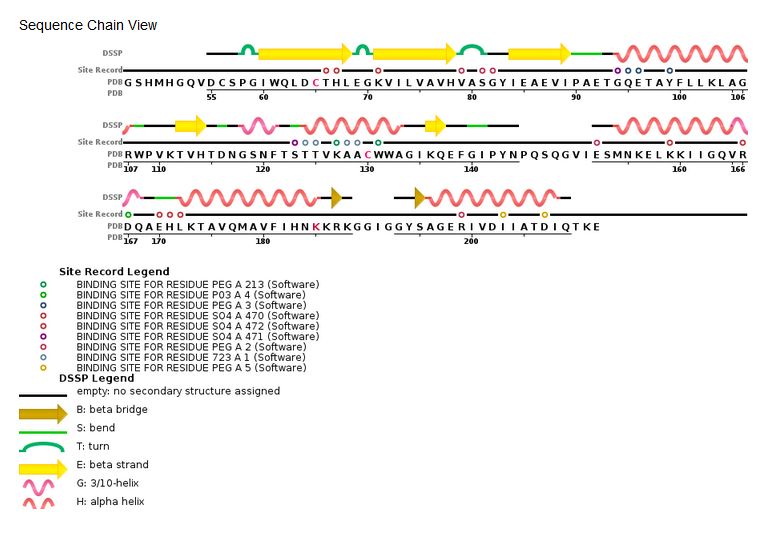
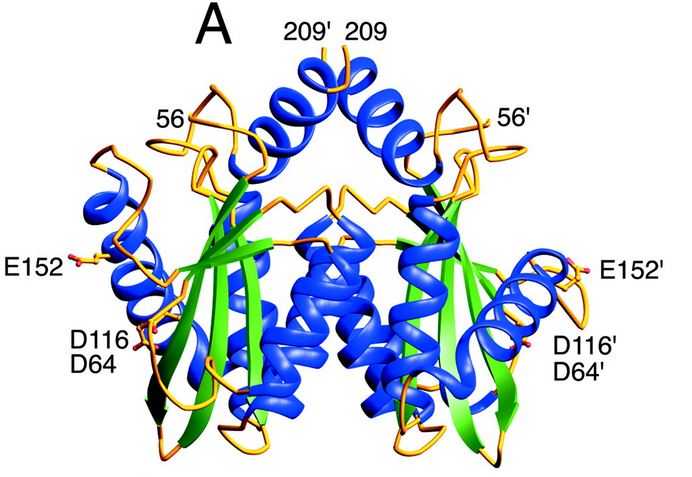
| - | ====Structure and role of the core domain in the integration==== | ||
'''The three domain of the HIV integrase:''' | '''The three domain of the HIV integrase:''' | ||
| Line 56: | Line 55: | ||
<table><tr><td colspan='2'>[[1bl3]] is a 3 chain structure with sequence from [http://en.wikipedia.org/wiki/9hiv1 9hiv1]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1BL3 OCA]. For a <b>guided tour on the structure components</b> use [http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=1BL3 FirstGlance]. <br> | <table><tr><td colspan='2'>[[1bl3]] is a 3 chain structure with sequence from [http://en.wikipedia.org/wiki/9hiv1 9hiv1]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1BL3 OCA]. For a <b>guided tour on the structure components</b> use [http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=1BL3 FirstGlance]. <br> | ||
</td></tr><tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat"><scene name='pdbligand=MG:MAGNESIUM+ION'>MG</scene></td></tr> | </td></tr><tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat"><scene name='pdbligand=MG:MAGNESIUM+ION'>MG</scene></td></tr> | ||
| + | |||
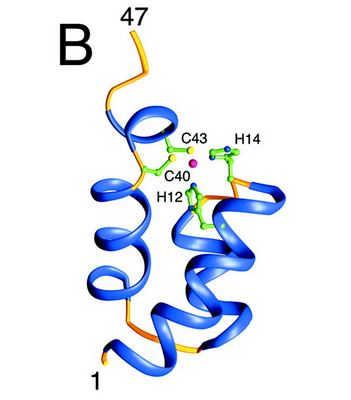
====Structure and role of the N ter domain==== | ====Structure and role of the N ter domain==== | ||
Revision as of 23:31, 10 January 2019
| This Sandbox is Reserved from 06/12/2018, through 30/06/2019 for use in the course "Structural Biology" taught by Bruno Kieffer at the University of Strasbourg, ESBS. This reservation includes Sandbox Reserved 1480 through Sandbox Reserved 1543. |
To get started:
More help: Help:Editing |
3lpt - HIV integrase
| |||||||||||