This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Frataxin
From Proteopedia
(Difference between revisions)
| Line 5: | Line 5: | ||
'''General Aspects''' | '''General Aspects''' | ||
| - | '''Frataxin''' is a protein capable of storing, releasing and detoxifying intracellular iron. In humans, a mutation in this protein can trigger the Friedreich's ataxia, a neurodegenerative disease caused due to incapacity to form iron-sulfur groups necessary to activate the mitochondrial enzyme involved in the electron transportation chain, aconitase. | + | '''Frataxin''' (FXN) is a protein capable of storing, releasing and detoxifying intracellular iron. In humans, a mutation in this protein can trigger the Friedreich's ataxia, a neurodegenerative disease caused due to incapacity to form iron-sulfur groups necessary to activate the mitochondrial enzyme involved in the electron transportation chain, aconitase. |
| - | It consists of a polymeric molecule that, | + | It consists of a polymeric molecule that, although capable of forming larger complexes (as the 24 subunit oligomer detected by electron microscopy), exerts its activity by association of three subunits, enough to form a central channel where the ferroxidation takes place. |
| - | In the following paragraphs, we describe the general features of its structure in the trimeric form as | + | In the following paragraphs, we describe the general features of its structure in the trimeric form as obtained by X-ray crystallography at 3Å resolution. The protein used was obtained from the Y37A yeast, which has a 40% sequence identity to the human frataxin. |
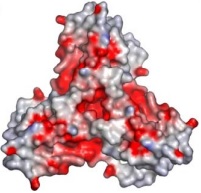
In the box at the right, it is possible to see its <scene name='78/788815/Spacefill_model/1'>general structure</scene> in a space-fill model, in which <font color='violet'><b>violet</b></font>, <font color='orangered'><b>orange</b></font> and <span style="color:aquamarine;background-color:darkgrey;font-weight:bold;">light-green</span> represent, each, a different monomer from the entire molecule. | In the box at the right, it is possible to see its <scene name='78/788815/Spacefill_model/1'>general structure</scene> in a space-fill model, in which <font color='violet'><b>violet</b></font>, <font color='orangered'><b>orange</b></font> and <span style="color:aquamarine;background-color:darkgrey;font-weight:bold;">light-green</span> represent, each, a different monomer from the entire molecule. | ||
| Line 18: | Line 18: | ||
'''General Secondary and Tertiary Structure Patterns''' | '''General Secondary and Tertiary Structure Patterns''' | ||
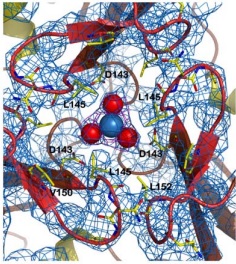
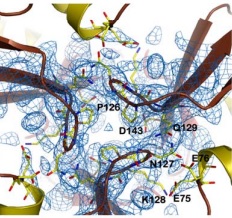
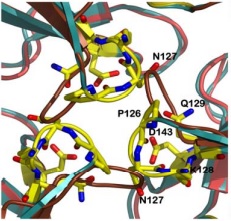
| - | The <scene name='78/788815/Alpha-helices_and_beta-sheets/1'>secondary structure</scene> is basically composed of two alpha-helices (in <font color='red'><b>red</b></font>) and a short helical segment (in <font color='magenta'><b>magenta</b></font>), and seven antiparallel | + | The <scene name='78/788815/Alpha-helices_and_beta-sheets/1'>secondary structure</scene> is basically composed of two alpha-helices (in <font color='red'><b>red</b></font>) and a short helical segment (in <font color='magenta'><b>magenta</b></font>), and seven antiparallel β-sheets, '''in each monomer''', represented here as six <span style="color:springgreen;background-color:darkgrey;font-weight:bold;">green setae</span> and one short region colored <span style="color:yellow;background-color:darkgrey;font-weight:bold;">yellow</span>. <font color='dodgerblue'><b>Turns</b></font> and <span style="color:white;background-color:darkseagreen;font-weight:bold;">coils</span> are also represented. |
You can also view the monomer isolated <scene name='78/788815/Isolated_monomer/1'>here</scene>. | You can also view the monomer isolated <scene name='78/788815/Isolated_monomer/1'>here</scene>. | ||
Revision as of 11:43, 15 January 2019
| |||||||||||
3D Structures of frataxin
Updated on 15-January-2019
References
KARLBERG, Tobias et al. The structures of frataxin oligomers reveal the mechanism for the delivery and detoxification of iron. Structure, v. 14, n. 10, p. 1535-1546, 2006.
Proteopedia Page Contributors and Editors (what is this?)
João Victor Paccini Coutinho, Michal Harel, Rebeca B. Candia