This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Aminopeptidase
From Proteopedia
(Difference between revisions)
| Line 22: | Line 22: | ||
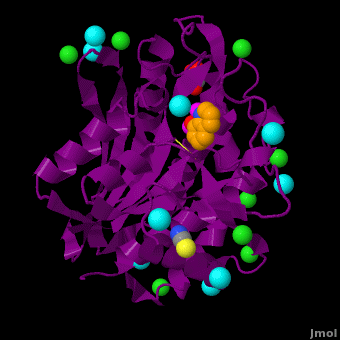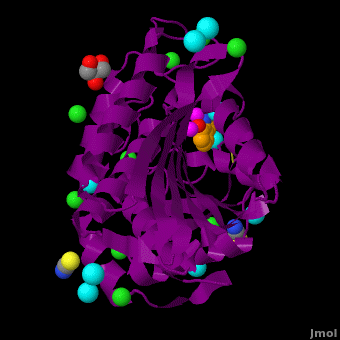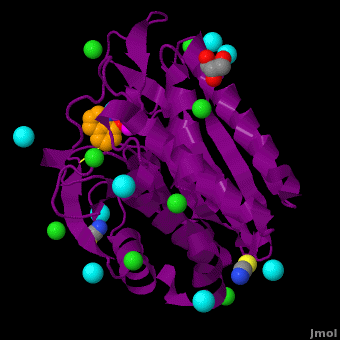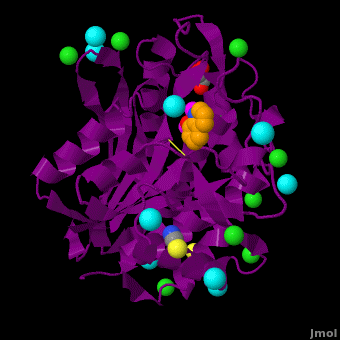
The <scene name='Aminopeptidase/Active_site/1'>active site</scene> of the enzyme contains two Zn2+ ions with His85 and Asp160 as ligands for one ion, and Glu132 and His247 as ligands for the second ion. Asp97 is a common ligand to both ions. What appears to be a phosphate anion is bound to both zinc atoms, replacing the water molecule/hydroxide ion normally found in this class of enzyme. See details of SGAP in [[Streptomyces griseus Aminopeptidase (SGAP)]]. | The <scene name='Aminopeptidase/Active_site/1'>active site</scene> of the enzyme contains two Zn2+ ions with His85 and Asp160 as ligands for one ion, and Glu132 and His247 as ligands for the second ion. Asp97 is a common ligand to both ions. What appears to be a phosphate anion is bound to both zinc atoms, replacing the water molecule/hydroxide ion normally found in this class of enzyme. See details of SGAP in [[Streptomyces griseus Aminopeptidase (SGAP)]]. | ||
| + | |||
| + | == 3D Structures of Aminopeptidase == | ||
| + | [[Aminopeptidase 3D structures]] | ||
| + | |||
</StructureSection> | </StructureSection> | ||
Revision as of 10:51, 13 March 2019
| |||||||||||
3D Structures of Aminopeptidase
Updated on 13-March-2019
Additional Resources
For additional information, see:
Amino Acid Synthesis & Metabolism
Streptomyces griseus Aminopeptidase (SGAP)
References
- ↑ Taylor A. Aminopeptidases: structure and function. FASEB J. 1993 Feb 1;7(2):290-8. PMID:8440407
- ↑ Hanaya K, Suetsugu M, Saijo S, Yamato I, Aoki S. Potent inhibition of dinuclear zinc(II) peptidase, an aminopeptidase from Aeromonas proteolytica, by 8-quinolinol derivatives: inhibitor design based on Zn(2+) fluorophores, kinetic, and X-ray crystallographic study. J Biol Inorg Chem. 2012 Feb 5. PMID:22311113 doi:10.1007/s00775-012-0873-4
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, David Canner, Joel L. Sussman, Eran Hodis