User:Asif Hossain/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 4: | Line 4: | ||
== Introduction == | == Introduction == | ||
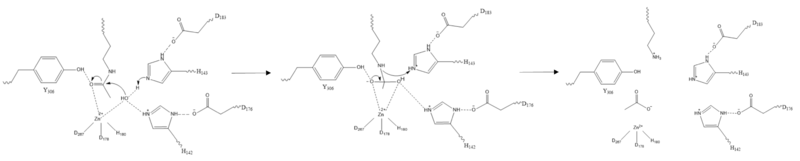
| - | Histone deacetylase 8 (HDAC8) is an enzyme that plays a role in controlling gene expression in ''Homo Sapiens''. Specifically, HDAC8 catalyzes the removal of an acetyl group off of the ε-amino-lysine sidechain of N-terminal core of [https://en.wikipedia.org/wiki/Histone histones].<ref name="Vannini, A., Volpari, C., Gallinari, P.">Vannini, A., Volpari, C., Gallinari, P., Jones, P., Mattu, M., Carfí, A., ... & Di Marco, S. (2007). Substrate binding to histone deacetylases as shown by the crystal structure of the HDAC8–substrate complex. EMBO reports, 8(9), 879-884. https://doi.org/10.1038/sj.embor.7401047 </ref> Histones consist of eight monomers to form an octomer complex. | + | Histone deacetylase 8 (HDAC8) is an enzyme that plays a role in controlling gene expression in ''Homo Sapiens''. Specifically, HDAC8 catalyzes the removal of an acetyl group off of the ε-amino-lysine sidechain of N-terminal core of [https://en.wikipedia.org/wiki/Histone histones].<ref name="Vannini, A., Volpari, C., Gallinari, P.">Vannini, A., Volpari, C., Gallinari, P., Jones, P., Mattu, M., Carfí, A., ... & Di Marco, S. (2007). Substrate binding to histone deacetylases as shown by the crystal structure of the HDAC8–substrate complex. EMBO reports, 8(9), 879-884. https://doi.org/10.1038/sj.embor.7401047 </ref> Histones consist of eight monomers to form an octomer complex. Each histone has a large positive charge. Since DNA is negatively charged, histones tightly interact with DNA. This prevents transcription factors from accessing DNA, thus decreasing gene expression. Chromatin remodeling by histone acetylation and/or deacetylation is an example of [https://en.wikipedia.org/wiki/Epigenetics epigenetic regulation]. [http://proteopedia.org/wiki/index.php/User:Nicholas_Bantz/Sandbox_1/ Histone acetlytransferase] (HAT1) catalyzes the addition of an acetyl group onto a histone. The lack of charge on the acetyl group prevents the interaction between DNA and histones. This allows transcription factors to access the DNA to increase gene expression. HDAC8 though reverses this reaction by catalyzing the removal of these acetyl groups by removing the acetate and the reclaimed positive charge on the lysine sidechain is able to interact with the negative charge on the DNA. As a result, DNA will bind more tightly to the histone protein, repressing transcription and gene expression. |
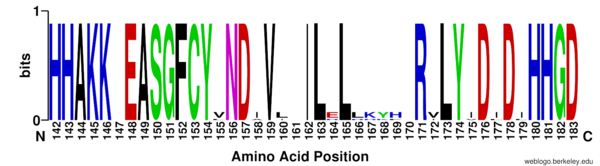
==Homology== | ==Homology== | ||
Revision as of 18:37, 9 April 2019
Histone Deacetylase 8 (HDAC 8)
| |||||||||||
References
- ↑ 1.0 1.1 1.2 1.3 1.4 Vannini, A., Volpari, C., Gallinari, P., Jones, P., Mattu, M., Carfí, A., ... & Di Marco, S. (2007). Substrate binding to histone deacetylases as shown by the crystal structure of the HDAC8–substrate complex. EMBO reports, 8(9), 879-884. https://doi.org/10.1038/sj.embor.7401047
- ↑ DesJarlais, R., & Tummino, P. J. (2016). Role of histone-modifying enzymes and their complexes in regulation of chromatin biology. Biochemistry, 55(11), 1584-1599. https://doi.org/10.1021/acs.biochem.5b01210
- ↑ 3.0 3.1 3.2 3.3 Somoza J, Skene R. Structural snapshots of human HDAC8 provide insights into the class I histone deacetylases. Structure, 12(7), 1325-1334.2004. https://doi.org/10.1016/j.str.2004.04.012
- ↑ Whitehead, L., Dobler, M. R., Radetich, B., Zhu, Y., Atadja, P. W., Claiborne, T., ... & Shao, W. (2011). Human HDAC isoform selectivity achieved via exploitation of the acetate release channel with structurally unique small molecule inhibitors. Bioorganic & medicinal chemistry, 19(15), 4626-4634. https://doi.org/10.1016/j.bmc.2011.06.030
- ↑ Seto, E., & Yoshida, M. (2014). Erasers of histone acetylation: the histone deacetylase enzymes. Cold Spring Harbor perspectives in biology, 6(4), a018713. https://doi.org/10.1101/cshperspect.a018713
- ↑ Eckschlager T, Plch, J, Stiborova M, Hrabeta J.Histone deacetylase inhibitors as anticancer drugs. International journal of molecular sciences, 18(7), 1414. 2017. https://dx.doi.org/10.3390%2Fijms18071414
- ↑ Vannini, A., Volpari, C., Filocamo, G., Casavola, E. C., Brunetti, M., Renzoni, D., ... & Steinkühler, C. (2004). Crystal structure of a eukaryotic zinc-dependent histone deacetylase, human HDAC8, complexed with a hydroxamic acid inhibitor. Proceedings of the National Academy of Sciences, 101(42), 15064-15069. https://dx.doi.org/10.1073%2Fpnas.0404603101