User:Asif Hossain/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 7: | Line 7: | ||
==HDAC Enzymes and Homology== | ==HDAC Enzymes and Homology== | ||
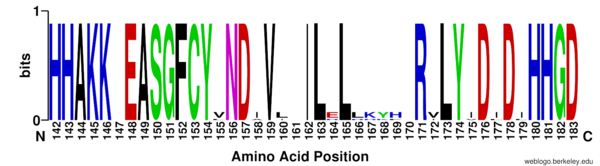
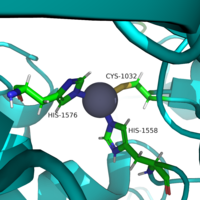
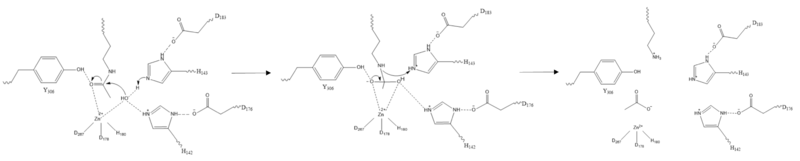
| - | There are four major classes of HDAC proteins (I,II, III, and IV). Other than the Class III “[https://en.wikipedia.org/wiki/Sirtuin sirtuins]” that utilize a NAD+ cofactor-dependent mechanism, all other HDAC classes use Zn<sup>2+</sup>-assisted catalysis through mechanisms reminiscent of a typical [https://en.wikipedia.org/wiki/Serine_protease serine protease].<ref name="DesJarlais, R., & Tummino, P. J.">DesJarlais, R., & Tummino, P. J. (2016). Role of histone-modifying enzymes and their complexes in regulation of chromatin biology. Biochemistry, 55(11), 1584-1599. https://doi.org/10.1021/acs.biochem.5b01210 </ref> While Classes I, II, and IV do have some major distinctions such as size of | + | There are four major classes of HDAC proteins (I,II, III, and IV). Other than the Class III “[https://en.wikipedia.org/wiki/Sirtuin sirtuins]” that utilize a NAD<sup>+</sup> cofactor-dependent mechanism, all other HDAC classes use Zn<sup>2+</sup>-assisted catalysis through mechanisms reminiscent of a typical [https://en.wikipedia.org/wiki/Serine_protease serine protease].<ref name="DesJarlais, R., & Tummino, P. J.">DesJarlais, R., & Tummino, P. J. (2016). Role of histone-modifying enzymes and their complexes in regulation of chromatin biology. Biochemistry, 55(11), 1584-1599. https://doi.org/10.1021/acs.biochem.5b01210 </ref> While Classes I, II, and IV do have some major distinctions such as size of peptide, they retain a great degree of homology both between and within classes, especially at catalytic sites. HDAC 8 is among the Class I HDACs alongside HDACs 1-3. It shares a high degree of conserved residues involved in the catalytic site, Zn-binding, and ligand binding pocket. (Figure 1) <ref name="Vannini, A., Volpari, C., Gallinari, P.">Vannini, A., Volpari, C., Gallinari, P., Jones, P., Mattu, M., Carfí, A., ... & Di Marco, S. (2007). Substrate binding to histone deacetylases as shown by the crystal structure of the HDAC8–substrate complex. EMBO reports, 8(9), 879-884. https://doi.org/10.1038/sj.embor.7401047 </ref> |
[[Image:Consensus.png|600px||center||thumb|Figure 1: Nearly all active site (Asp183/H143, Asp176/His142 shown) zinc binding (Asp178 and His180 shown), and binding pocket residues (Gly151 and Phe 152 shown) are conserved across homologous sequences amongst Class I HDACs. ]] | [[Image:Consensus.png|600px||center||thumb|Figure 1: Nearly all active site (Asp183/H143, Asp176/His142 shown) zinc binding (Asp178 and His180 shown), and binding pocket residues (Gly151 and Phe 152 shown) are conserved across homologous sequences amongst Class I HDACs. ]] | ||
Revision as of 18:42, 9 April 2019
Histone Deacetylase 8 (HDAC 8)
| |||||||||||
References
- ↑ 1.0 1.1 1.2 1.3 1.4 Vannini, A., Volpari, C., Gallinari, P., Jones, P., Mattu, M., Carfí, A., ... & Di Marco, S. (2007). Substrate binding to histone deacetylases as shown by the crystal structure of the HDAC8–substrate complex. EMBO reports, 8(9), 879-884. https://doi.org/10.1038/sj.embor.7401047
- ↑ DesJarlais, R., & Tummino, P. J. (2016). Role of histone-modifying enzymes and their complexes in regulation of chromatin biology. Biochemistry, 55(11), 1584-1599. https://doi.org/10.1021/acs.biochem.5b01210
- ↑ 3.0 3.1 3.2 3.3 Somoza J, Skene R. Structural snapshots of human HDAC8 provide insights into the class I histone deacetylases. Structure, 12(7), 1325-1334.2004. https://doi.org/10.1016/j.str.2004.04.012
- ↑ Whitehead, L., Dobler, M. R., Radetich, B., Zhu, Y., Atadja, P. W., Claiborne, T., ... & Shao, W. (2011). Human HDAC isoform selectivity achieved via exploitation of the acetate release channel with structurally unique small molecule inhibitors. Bioorganic & medicinal chemistry, 19(15), 4626-4634. https://doi.org/10.1016/j.bmc.2011.06.030
- ↑ Seto, E., & Yoshida, M. (2014). Erasers of histone acetylation: the histone deacetylase enzymes. Cold Spring Harbor perspectives in biology, 6(4), a018713. https://doi.org/10.1101/cshperspect.a018713
- ↑ Eckschlager T, Plch, J, Stiborova M, Hrabeta J.Histone deacetylase inhibitors as anticancer drugs. International journal of molecular sciences, 18(7), 1414. 2017. https://dx.doi.org/10.3390%2Fijms18071414
- ↑ Vannini, A., Volpari, C., Filocamo, G., Casavola, E. C., Brunetti, M., Renzoni, D., ... & Steinkühler, C. (2004). Crystal structure of a eukaryotic zinc-dependent histone deacetylase, human HDAC8, complexed with a hydroxamic acid inhibitor. Proceedings of the National Academy of Sciences, 101(42), 15064-15069. https://dx.doi.org/10.1073%2Fpnas.0404603101