This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Madeleine Wilson/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
| - | =Lysine | + | =Histone Lysine Methyltransferase: Gene Activator= |
<StructureSection load='1O9S' size='350' frame='true' side='right' caption='Lysine Methyl Transferase' scene=’C_terminal_domain’> | <StructureSection load='1O9S' size='350' frame='true' side='right' caption='Lysine Methyl Transferase' scene=’C_terminal_domain’> | ||
| Line 6: | Line 6: | ||
===Histone Methylation=== | ===Histone Methylation=== | ||
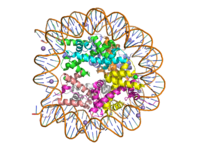
| - | [[Image:human_nucleosome_ray_trace.png|200 px| right| thumb|Human nucleosome]] | + | [[Image:human_nucleosome_ray_trace.png|200 px| right| thumb|Human nucleosome, pbd code: 5y0c]] |
Histone proteins aid in the packing of DNA for the purpose of compacting the genome in the nucleus of the cell and regulating physical accessibility of genes for transcription. The protein itself is an octamer made of heterodimer core proteins H2a, H2b, H3, and H4, with H1 and H5 acting as linker proteins. About 145-157 base pairs wind around a histone core protein. <ref name="DesJarlais">PMID: 26745824</ref> Modifications to histone core proteins can affect the accessibility of genes in the genome and their ability to be transcribed. Some of these modifications include methylation/demethylation, acetylation/deacetylation, and ubiquitination/deubiquitination. <ref name="Lun">DOI: 10.1016/j.apsb.2013.04.007</ref> | Histone proteins aid in the packing of DNA for the purpose of compacting the genome in the nucleus of the cell and regulating physical accessibility of genes for transcription. The protein itself is an octamer made of heterodimer core proteins H2a, H2b, H3, and H4, with H1 and H5 acting as linker proteins. About 145-157 base pairs wind around a histone core protein. <ref name="DesJarlais">PMID: 26745824</ref> Modifications to histone core proteins can affect the accessibility of genes in the genome and their ability to be transcribed. Some of these modifications include methylation/demethylation, acetylation/deacetylation, and ubiquitination/deubiquitination. <ref name="Lun">DOI: 10.1016/j.apsb.2013.04.007</ref> | ||
Revision as of 18:40, 16 April 2019
Histone Lysine Methyltransferase: Gene Activator
| |||||||||||
References
- ↑ DesJarlais R, Tummino PJ. Role of Histone-Modifying Enzymes and Their Complexes in Regulation of Chromatin Biology. Biochemistry. 2016 Mar 22;55(11):1584-99. doi: 10.1021/acs.biochem.5b01210. Epub , 2016 Jan 26. PMID:26745824 doi:http://dx.doi.org/10.1021/acs.biochem.5b01210
- ↑ 2.0 2.1 doi: https://dx.doi.org/10.1016/j.apsb.2013.04.007
- ↑ 3.0 3.1 Dong X, Weng Z. The correlation between histone modifications and gene expression. Epigenomics. 2013 Apr;5(2):113-6. doi: 10.2217/epi.13.13. PMID:23566087 doi:http://dx.doi.org/10.2217/epi.13.13
- ↑ 4.0 4.1 4.2 4.3 Xiao B, Jing C, Wilson JR, Walker PA, Vasisht N, Kelly G, Howell S, Taylor IA, Blackburn GM, Gamblin SJ. Structure and catalytic mechanism of the human histone methyltransferase SET7/9. Nature. 2003 Feb 6;421(6923):652-6. Epub 2003 Jan 22. PMID:12540855 doi:10.1038/nature01378
- ↑ Del Rizzo PA, Trievel RC. Substrate and product specificities of SET domain methyltransferases. Epigenetics. 2011 Sep 1;6(9):1059-67. doi: 10.4161/epi.6.9.16069. Epub 2011 Sep, 1. PMID:21847010 doi:http://dx.doi.org/10.4161/epi.6.9.16069
- ↑ Schluckebier G, Kozak M, Bleimling N, Weinhold E, Saenger W. Differential binding of S-adenosylmethionine S-adenosylhomocysteine and Sinefungin to the adenine-specific DNA methyltransferase M.TaqI. J Mol Biol. 1997 Jan 10;265(1):56-67. PMID:8995524 doi:http://dx.doi.org/10.1006/jmbi.1996.0711
- ↑ Tamura R, Doi S, Nakashima A, Sasaki K, Maeda K, Ueno T, Masaki T. Inhibition of the H3K4 methyltransferase SET7/9 ameliorates peritoneal fibrosis. PLoS One. 2018 May 3;13(5):e0196844. doi: 10.1371/journal.pone.0196844., eCollection 2018. PMID:29723250 doi:http://dx.doi.org/10.1371/journal.pone.0196844
Student Contributors
Lauryn Padgett, Alexandra Pentala, Madeleine Wilson