This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Caitlin Marie Gaich/Sandbox1
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
=Histone Acetyltransferase HAT1/HAT2 Complex, ''Saccharomyces cerevisiae''= | =Histone Acetyltransferase HAT1/HAT2 Complex, ''Saccharomyces cerevisiae''= | ||
<StructureSection load='4PSW' size='350' frame='true' side='right' caption='HAT1/HAT2 Complex pdb: 4PSW' scene='81/811717/Overview/1'> | <StructureSection load='4PSW' size='350' frame='true' side='right' caption='HAT1/HAT2 Complex pdb: 4PSW' scene='81/811717/Overview/1'> | ||
| - | + | =Histones= | |
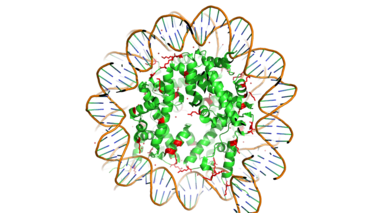
[https://en.wikipedia.org/wiki/Histone Histones] are proteins found in the nucleus that are the key building blocks of [https://en.wikipedia.org/wiki/Chromatin chromatin] and are essential for proper DNA packaging and [https://en.wikipedia.org/wiki/Transcription_(biology) transcription]. In the first step of [https://www.hhmi.org/biointeractive/how-dna-packaged DNA packaging], two copies of the four core histone proteins (H1A, H2A, H3, and H4) form an [https://en.wikipedia.org/wiki/Histone_octamer octamer] in which DNA directly interacts with and wraps around, forming the [https://en.wikipedia.org/wiki/Nucleosome nucleosome]. 20-24% of residues making up the histone octamer are arginine and lysine, causing a net positive charge, especially at the outer surfaces of the histone core where negatively-charged DNA is bound. <ref> Watson, J D, et al. Molecular Biology of the Gene (Seventh Edition). (2014) Boston, MA: Benjamin-Cummings Publishing Company. </ref> It is those positively charged tails of the histone core that are often subject to post-translational modifications that play important roles in replication, transcription, heterochromatin maintenance, and DNA repair. | [https://en.wikipedia.org/wiki/Histone Histones] are proteins found in the nucleus that are the key building blocks of [https://en.wikipedia.org/wiki/Chromatin chromatin] and are essential for proper DNA packaging and [https://en.wikipedia.org/wiki/Transcription_(biology) transcription]. In the first step of [https://www.hhmi.org/biointeractive/how-dna-packaged DNA packaging], two copies of the four core histone proteins (H1A, H2A, H3, and H4) form an [https://en.wikipedia.org/wiki/Histone_octamer octamer] in which DNA directly interacts with and wraps around, forming the [https://en.wikipedia.org/wiki/Nucleosome nucleosome]. 20-24% of residues making up the histone octamer are arginine and lysine, causing a net positive charge, especially at the outer surfaces of the histone core where negatively-charged DNA is bound. <ref> Watson, J D, et al. Molecular Biology of the Gene (Seventh Edition). (2014) Boston, MA: Benjamin-Cummings Publishing Company. </ref> It is those positively charged tails of the histone core that are often subject to post-translational modifications that play important roles in replication, transcription, heterochromatin maintenance, and DNA repair. | ||
[[Image:Histone_w_DNA.png|380 px|right|thumb|Figure 1. Histone Core w/ DNA Bound]] | [[Image:Histone_w_DNA.png|380 px|right|thumb|Figure 1. Histone Core w/ DNA Bound]] | ||
| - | + | =Histone Modification= | |
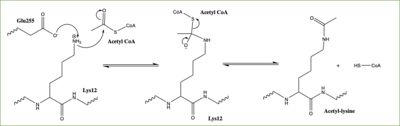
Histones can be modified in a variety of ways, including: methylation, demethylation, acetylation, deacetylation and many others, all leading to either the condensation or relaxation of DNA and as a consequence turning on or off DNA transcription. Histone acetylation is a common histone modification that involves the transfer of an acetyl moiety from Acetyl Coenzyme A (AcCoA) to an ε-amino group of the target lysine residue on a histone. This reaction is catalyzed by the histone acetyltransferase (HAT) enzyme families. The specific histone acetylation modification is an important [https://en.wikipedia.org/wiki/Epigenetics epigenetic] marker. It plays a role in RNA synthesis and there a known correlation between gene activity and histone acetylation. Any misregulations of the HAT enzyme can possibly lead to cancer, cardiovascular disease, and HIV. | Histones can be modified in a variety of ways, including: methylation, demethylation, acetylation, deacetylation and many others, all leading to either the condensation or relaxation of DNA and as a consequence turning on or off DNA transcription. Histone acetylation is a common histone modification that involves the transfer of an acetyl moiety from Acetyl Coenzyme A (AcCoA) to an ε-amino group of the target lysine residue on a histone. This reaction is catalyzed by the histone acetyltransferase (HAT) enzyme families. The specific histone acetylation modification is an important [https://en.wikipedia.org/wiki/Epigenetics epigenetic] marker. It plays a role in RNA synthesis and there a known correlation between gene activity and histone acetylation. Any misregulations of the HAT enzyme can possibly lead to cancer, cardiovascular disease, and HIV. | ||
Revision as of 01:24, 25 April 2019
Histone Acetyltransferase HAT1/HAT2 Complex, Saccharomyces cerevisiae
| |||||||||||