This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Lauryn Padgett/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 12: | Line 12: | ||
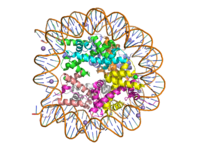
==Lysine Methyltransferase (KMT) Structure== | ==Lysine Methyltransferase (KMT) Structure== | ||
| - | The structure of human histone methyltransferase SET7/9 was crystallized using x-ray diffraction at 1.75Å in its product form <ref name="Xiao" />. SET7/9 was crystallized with its cofactor was in its unmethylated form, <scene name='81/811086/Sah/1'>S-adenosyl-L-homocysteine</scene> (SAH), and with its <scene name='81/811086/10resproduct/1'>product</scene>, a ten residue peptide with a methylated lysine at residue 4. | + | The structure of human histone methyltransferase SET7/9 was crystallized using x-ray diffraction at 1.75Å in its product form <ref name="Xiao" />. SET7/9 was crystallized with its cofactor was in its unmethylated form, <scene name='81/811086/Sah/1'>S-adenosyl-L-homocysteine</scene> (SAH), and with its <scene name='81/811086/10resproduct/1'>product</scene>, a ten residue peptide with a methylated lysine at residue 4. The PDB code for the crystallized SET7/9 structure is 1O9S. |
===Overall Secondary Structure=== | ===Overall Secondary Structure=== | ||
| - | Due to the composition of its [https://en.wikipedia.org/wiki/Protein_secondary_structure secondary structure], KMT is an alpha-beta protein <ref name="Xiao" />. The helical composition includes 3 <scene name='81/811086/Alpha_helices/4'>alpha helices</scene>, with two residing in the SET domain and one in the C-terminal domain. The alpha helices in the SET domain are two turns while the C-terminal helix is | + | Due to the composition of its [https://en.wikipedia.org/wiki/Protein_secondary_structure secondary structure], KMT is an alpha-beta protein <ref name="Xiao" />. The helical composition includes 3 <scene name='81/811086/Alpha_helices/4'>alpha helices</scene>, with two residing in the SET domain and one in the C-terminal domain. The alpha helices in the SET domain are two turns while the C-terminal alpha helix is the largest with 4 turns. There are also 2 <scene name='81/811086/3-10_helices/4'>3-10 helices</scene> in the SET domain which are each one turn. There are 21 total <scene name='81/811086/Beta_sheets/3'>beta strands</scene> which reside in both the N-terminal domain and the SET domain. The beta strands are primarily anti-parallel and multiple antiparallel strands are connected by Type 1 and Type 2 <scene name='81/811086/Beta_turns/3'>beta turns</scene>. |
===The Active Site=== | ===The Active Site=== | ||
| Line 32: | Line 32: | ||
==Inhibitors== | ==Inhibitors== | ||
| + | |||
| + | SET7/9’s structure and function has been studied extensively because of its role in transcription <ref name="Takemoto">PMID:27088648</ref>. In the past few years it has been identified to methylate genes involved in multiple diseases; making it a potential candidate for drug inhibition <ref name="Takemoto"> <ref name="Schluckebier"> <ref name="Tamura">. Two compounds that have been found to inhibit SET7/9 in certain cells in vitro are Sinefungin and Cyproheptadine. Each inhibitor acts on the catalytic center of SET7/9, however their mechanisms of inhibition and possible medical relevancies differ greatly. | ||
===Sinefungin=== | ===Sinefungin=== | ||
| Line 37: | Line 39: | ||
<scene name='81/811086/Betahairincypr/1'>Normalhairpin</scene> | <scene name='81/811086/Betahairincypr/1'>Normalhairpin</scene> | ||
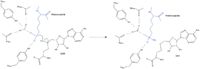
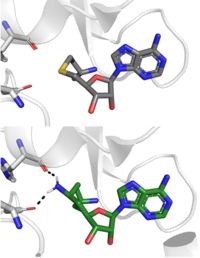
| - | [[Image: SinSAH.jpg|200 px| right| thumb|SAH (grey) and Sinefungin (green) in the peptide binding pocket. The nitrogen group of sinefungin makes 2 double bonds to the main chain carbonyls of | + | [[Image: SinSAH.jpg|200 px| right| thumb|SAH (grey) and Sinefungin (green) in the peptide binding pocket. The nitrogen group of sinefungin makes 2 double bonds to the main chain carbonyls of Arg265 and His293. Sinefungin was created using PDB: 1O9S and mutating the sulfur of SAH]] |
Sinefungin has been used experimentally to inhibit the SET 7/9 protein on [https://www.sciencedirect.com/topics/medicine-and-dentistry/peritoneal-fibrosis peritoneal fibrosis] in mice and in human peritoneal mesothelial cells <ref name=”Tamura”>PMID:29723250</ref>. SET 7/9 is involved in peritoneal fibrosis because it mono-methylates H3K4, which activates the transcription of fibrosis related genes. The administration of Sinefungin to mice in vitro resulted in decreased levels of methylated H3K4 (H3K4me1) protein, as well as suppressed peritoneal cell density and thickening. The decreased levels of H3K4me1 suggest that the methylation of H3K4 was inhibited by Sinefungin, as well as that inhibiting SET7/9 ameliorates peritoneal fibrosis. | Sinefungin has been used experimentally to inhibit the SET 7/9 protein on [https://www.sciencedirect.com/topics/medicine-and-dentistry/peritoneal-fibrosis peritoneal fibrosis] in mice and in human peritoneal mesothelial cells <ref name=”Tamura”>PMID:29723250</ref>. SET 7/9 is involved in peritoneal fibrosis because it mono-methylates H3K4, which activates the transcription of fibrosis related genes. The administration of Sinefungin to mice in vitro resulted in decreased levels of methylated H3K4 (H3K4me1) protein, as well as suppressed peritoneal cell density and thickening. The decreased levels of H3K4me1 suggest that the methylation of H3K4 was inhibited by Sinefungin, as well as that inhibiting SET7/9 ameliorates peritoneal fibrosis. | ||
| Line 43: | Line 45: | ||
===Cyproheptadine=== | ===Cyproheptadine=== | ||
| - | Another inhibitor of SET 7/9 is <scene name='81/811086/Cyproheptadine/3'>cyproheptadine</scene>, a clinically-approved anti-allergy drug that was originally developed as a serotonin and histimine <ref name="Takemoto" | + | Another inhibitor of SET 7/9 is <scene name='81/811086/Cyproheptadine/3'>cyproheptadine</scene>, a clinically-approved anti-allergy drug that was originally developed as a serotonin and histimine <ref name="Takemoto">. The cyproheptadine-SET 7/9 complex was crystallized via x-ray diffraction at 2.005 Å with methylated cofactor SAM and with cydroheptadine. Unlike Sinefungin, it is a traditional competitor and competitive with the peptide substrates as it binds to the peptide-binding site. When cyproheptadine binds to the substrate site, the nitrogen of the [https://www.koeichem.com/en/product/index.php/item?cell003=Amines&cell004=Piperidine+derivatives&page=8&name=N-Methylpiperidine&id=116&label=1 methylpiperdine] ring of cyproheptadine forms a hydrogen bond with Thr286 as well as hydrophobic and interactions with the residues surrounding its binding site. The binding of cyproheptadine to the catalytic site causes conformational changes of residue Tyr337, an important residue for the formation of the lysine access channel. This movement subsequently causes a conformational change of the beta hairpin, <scene name='81/811086/Betahairpincyp/2'>residues 337-349</scene> and ultimately generates a large hole adjacent to the lysine access channel, as well as a shift of the C-terminal helix. |
With the revelation of its inhibitory effects on SET7/9, cyproheptadine was used in vitro to treat breast cancer cells ([https://en.wikipedia.org/wiki/MCF-7 MCF-7] cells). SET 7/9's non-histone activities include the methylation of the estrogen receptor α (ERα), a nuclear receptor and a transcription factor responsible for estrogen-responsive gene regulation. The expression and transcriptional activity of ERα is involved in the carcinogenesis of 70% of breast cancers, making it a major target for hormone therapy. Researchers found that treating the MCF7 cells with cyproheptadine decreased ERα's expression and transcriptional activity which therefore inhibited the estrogen-dependent cell growth. These findings suggest that cyproheptadine could possibly be repurposed to breast cancer therapy in the future. | With the revelation of its inhibitory effects on SET7/9, cyproheptadine was used in vitro to treat breast cancer cells ([https://en.wikipedia.org/wiki/MCF-7 MCF-7] cells). SET 7/9's non-histone activities include the methylation of the estrogen receptor α (ERα), a nuclear receptor and a transcription factor responsible for estrogen-responsive gene regulation. The expression and transcriptional activity of ERα is involved in the carcinogenesis of 70% of breast cancers, making it a major target for hormone therapy. Researchers found that treating the MCF7 cells with cyproheptadine decreased ERα's expression and transcriptional activity which therefore inhibited the estrogen-dependent cell growth. These findings suggest that cyproheptadine could possibly be repurposed to breast cancer therapy in the future. | ||
Revision as of 21:29, 26 April 2019
Histone Lysine Methyltransferase: Gene Activator
| |||||||||||
References
- ↑ DesJarlais R, Tummino PJ. Role of Histone-Modifying Enzymes and Their Complexes in Regulation of Chromatin Biology. Biochemistry. 2016 Mar 22;55(11):1584-99. doi: 10.1021/acs.biochem.5b01210. Epub , 2016 Jan 26. PMID:26745824 doi:http://dx.doi.org/10.1021/acs.biochem.5b01210
- ↑ 2.0 2.1 doi: https://dx.doi.org/10.1016/j.apsb.2013.04.007
- ↑ 3.0 3.1 Dong X, Weng Z. The correlation between histone modifications and gene expression. Epigenomics. 2013 Apr;5(2):113-6. doi: 10.2217/epi.13.13. PMID:23566087 doi:http://dx.doi.org/10.2217/epi.13.13
- ↑ 4.0 4.1 Del Rizzo PA, Trievel RC. Substrate and product specificities of SET domain methyltransferases. Epigenetics. 2011 Sep 1;6(9):1059-67. doi: 10.4161/epi.6.9.16069. Epub 2011 Sep, 1. PMID:21847010 doi:http://dx.doi.org/10.4161/epi.6.9.16069
- ↑ 5.0 5.1 5.2 5.3 5.4 5.5 5.6 5.7 Xiao B, Jing C, Wilson JR, Walker PA, Vasisht N, Kelly G, Howell S, Taylor IA, Blackburn GM, Gamblin SJ. Structure and catalytic mechanism of the human histone methyltransferase SET7/9. Nature. 2003 Feb 6;421(6923):652-6. Epub 2003 Jan 22. PMID:12540855 doi:10.1038/nature01378
- ↑ 6.0 6.1 6.2 Takemoto Y, Ito A, Niwa H, Okamura M, Fujiwara T, Hirano T, Handa N, Umehara T, Sonoda T, Ogawa K, Tariq M, Nishino N, Dan S, Kagechika H, Yamori T, Yokoyama S, Yoshida M. Identification of Cyproheptadine as an Inhibitor of SET Domain Containing Lysine Methyltransferase 7/9 (Set7/9) That Regulates Estrogen-Dependent Transcription. J Med Chem. 2016 Apr 28;59(8):3650-60. doi: 10.1021/acs.jmedchem.5b01732. Epub, 2016 Apr 18. PMID:27088648 doi:http://dx.doi.org/10.1021/acs.jmedchem.5b01732
- ↑ Tamura R, Doi S, Nakashima A, Sasaki K, Maeda K, Ueno T, Masaki T. Inhibition of the H3K4 methyltransferase SET7/9 ameliorates peritoneal fibrosis. PLoS One. 2018 May 3;13(5):e0196844. doi: 10.1371/journal.pone.0196844., eCollection 2018. PMID:29723250 doi:http://dx.doi.org/10.1371/journal.pone.0196844
Student Contributors
Lauryn Padgett, Alexandra Pentala, Madeleine Wilson