This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Penicillin-binding protein
From Proteopedia
(Difference between revisions)
| Line 10: | Line 10: | ||
== Structural highlights == | == Structural highlights == | ||
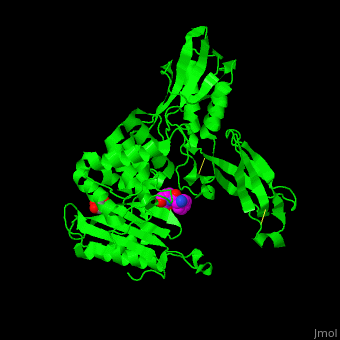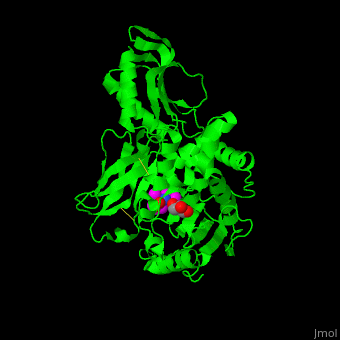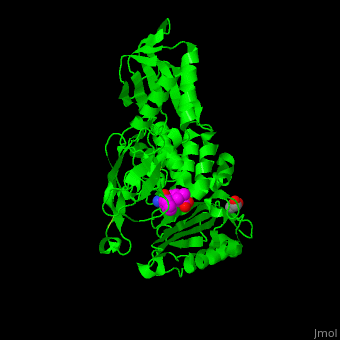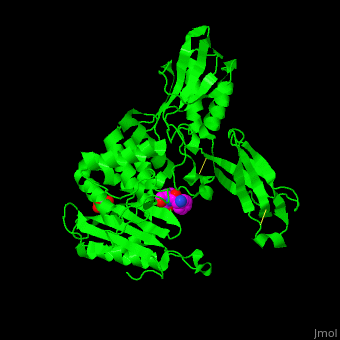
| - | ''E. coli'' PBP structure shows a distinct <scene name='47/478544/Cv/ | + | ''E. coli'' PBP structure shows a distinct <scene name='47/478544/Cv/5'>3 domain structures</scene>. The <scene name='47/478544/Cv/6'>active site</scene> contains the <scene name='47/478544/Cv/7'>covalently bond between Ser62 and antibiotic ampicillin</scene><ref>PMID:16411754</ref>. Water molecules are shown as red spheres. |
</StructureSection> | </StructureSection> | ||
==3D structures of penicillin-binding protein== | ==3D structures of penicillin-binding protein== | ||
Revision as of 11:49, 30 July 2019
| |||||||||||
3D structures of penicillin-binding protein
Updated on 30-July-2019
References
- ↑ Spratt BG. Distinct penicillin binding proteins involved in the division, elongation, and shape of Escherichia coli K12. Proc Natl Acad Sci U S A. 1975 Aug;72(8):2999-3003. PMID:1103132
- ↑ Beadle BM, Nicholas RA, Shoichet BK. Interaction energies between beta-lactam antibiotics and E. coli penicillin-binding protein 5 by reversible thermal denaturation. Protein Sci. 2001 Jun;10(6):1254-9. PMID:11369864 doi:http://dx.doi.org/10.1110/ps.52001
- ↑ Kishida H, Unzai S, Roper DI, Lloyd A, Park SY, Tame JR. Crystal structure of penicillin binding protein 4 (dacB) from Escherichia coli, both in the native form and covalently linked to various antibiotics. Biochemistry. 2006 Jan 24;45(3):783-92. PMID:16411754 doi:10.1021/bi051533t