This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Hydroxylase
From Proteopedia
(Difference between revisions)
| Line 41: | Line 41: | ||
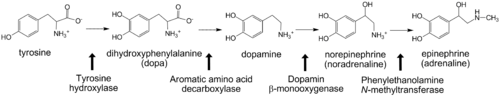
Deficiency in tyrosine hydroxylase has been observed in dopa responsive dystonia and juvenile parkinsonism. A reduced 3,4-dihydroxyphenylalanine production is also observed in Parkinson’s disease<ref>PMID: 11849022</ref>. | Deficiency in tyrosine hydroxylase has been observed in dopa responsive dystonia and juvenile parkinsonism. A reduced 3,4-dihydroxyphenylalanine production is also observed in Parkinson’s disease<ref>PMID: 11849022</ref>. | ||
| + | |||
| + | ==3D structures of hydroxylase== | ||
| + | [[Hydroxylases 3D structures]] | ||
| + | |||
</StructureSection> | </StructureSection> | ||
==3D structures of hydroxylases== | ==3D structures of hydroxylases== | ||
| Line 48: | Line 52: | ||
*Phenylalanine hydroxylase | *Phenylalanine hydroxylase | ||
| - | **[[ | + | **[[6n1k]] – hPAH (mutant) – human<br /> |
| + | **[[6hyc]] – hPAH + cofactor<br /> | ||
**[[1j8t]], [[1pah]], [[2pah]] - hPAH catalytic domain<br /> | **[[1j8t]], [[1pah]], [[2pah]] - hPAH catalytic domain<br /> | ||
| + | **[[1tdw]] – hPAH catalytic domain (mutant) <br /> | ||
| + | **[[6hpo]] – hPAH catalytic domain + Fe <br /> | ||
**[[3pah]], [[4pah]] – hPAH catalytic domain + adrenaline inhibitor<br /> | **[[3pah]], [[4pah]] – hPAH catalytic domain + adrenaline inhibitor<br /> | ||
**[[5pah]], [[6pah]] - hPAH catalytic domain + dopamine inhibitor<br /> | **[[5pah]], [[6pah]] - hPAH catalytic domain + dopamine inhibitor<br /> | ||
**[[1j8u]] - hPAH catalytic domain + cofactor<br /> | **[[1j8u]] - hPAH catalytic domain + cofactor<br /> | ||
| - | **[[1dmw]] - hPAH + cofactor<br /> | + | **[[1dmw]] - hPAH catalytic domain + cofactor<br /> |
| - | **[[1tg2]], [[1lrm]] - hPAH (mutant) + cofactor<br /> | + | **[[6hpo]] - hPAH catalytic domain + cofactor + Fe<br /> |
| + | **[[1tg2]], [[1lrm]] - hPAH catalytic domain (mutant) + cofactor<br /> | ||
**[[1mmk]], [[1mmt]], [[1kw0]] - hPAH catalytic domain + cofactor + substrate<br /> | **[[1mmk]], [[1mmt]], [[1kw0]] - hPAH catalytic domain + cofactor + substrate<br /> | ||
**[[4anp]] - hPAH catalytic domain + pyrimidine derivative<br /> | **[[4anp]] - hPAH catalytic domain + pyrimidine derivative<br /> | ||
**[[5fii]], [[5fgi]] - hPAH residues 19-118 + phenylalanine<br /> | **[[5fii]], [[5fgi]] - hPAH residues 19-118 + phenylalanine<br /> | ||
| + | **[[1phz]], [[2phm]], [[5den]], [[5fgj]], [[5den]] – rPAH - rat<br /> | ||
| + | **[[5egq]] - rPAH (mutant) <br /> | ||
**[[5jk6]] - smPAH + Fe – slime mold<br /> | **[[5jk6]] - smPAH + Fe – slime mold<br /> | ||
**[[5jk5]] - smPAH + cofactor + Fe <br /> | **[[5jk5]] - smPAH + cofactor + Fe <br /> | ||
| Line 105: | Line 115: | ||
**[[1k0l]] – PaPHBH (mutant) + FAD <br /> | **[[1k0l]] – PaPHBH (mutant) + FAD <br /> | ||
**[[6dll]] – PfPHBH + FAD – ''Pseudomonas putida''<br /> | **[[6dll]] – PfPHBH + FAD – ''Pseudomonas putida''<br /> | ||
| + | **[[2dki]] – CtPHBH + FAD – ''Comamonas testosteroni''<br /> | ||
| + | **[[2dkh]] – CtPHBH + FAD + hydroxy-benzoate <br /> | ||
*Apartyl/asparaginyl β-hydroxylase | *Apartyl/asparaginyl β-hydroxylase | ||
| Line 116: | Line 128: | ||
*Alpha-ketoglutarate-dependent L-arginine hydroxylase | *Alpha-ketoglutarate-dependent L-arginine hydroxylase | ||
| - | **[[6dax]], [[6alm]], [[6aln]] – SvAAH + arginine + oxoglutarate + Fe – ''Streptomyces vinaceus''<br /> | + | **[[6dax]], [[6alm]], [[6aln]], [[6mp8]] – SvAAH + arginine + oxoglutarate + Fe – ''Streptomyces vinaceus''<br /> |
**[[6daz]], [[6alo]], [[6alp]], [[6alq]] – SvAAH + arginine + succinate + Fe <br /> | **[[6daz]], [[6alo]], [[6alp]], [[6alq]] – SvAAH + arginine + succinate + Fe <br /> | ||
**[[6db2]], [[6alr]] – SvAAH + homoarginine + succinate + V <br /> | **[[6db2]], [[6alr]] – SvAAH + homoarginine + succinate + V <br /> | ||
| + | **[[6mp9]] – SvAAH + guanidinopentanoate + succinate + Fe <br /> | ||
*PLP-dependent L-arginine hydroxylase | *PLP-dependent L-arginine hydroxylase | ||
| Line 125: | Line 138: | ||
**[[5dj1]] – SwAAH + PLP + Mg<br /> | **[[5dj1]] – SwAAH + PLP + Mg<br /> | ||
**[[5dj3]] – SwAAH + PLP-arginine + Mg<br /> | **[[5dj3]] – SwAAH + PLP-arginine + Mg<br /> | ||
| + | **[[5bk7]] – SwAAH (mutant) + PLP-arginine <br /> | ||
| + | **[[6c8t]], [[6c92]] – SwAAH + PLP-arginine + PLP derivative<br /> | ||
*Leucine hydroxylase | *Leucine hydroxylase | ||
| Line 131: | Line 146: | ||
**[[5nci]] – StAAH + leucine derivative + oxoglutarate + Co <br /> | **[[5nci]] – StAAH + leucine derivative + oxoglutarate + Co <br /> | ||
**[[5ncj]] – StAAH + leucine derivative + succinate + Mn <br /> | **[[5ncj]] – StAAH + leucine derivative + succinate + Mn <br /> | ||
| + | |||
| + | *L-arginine b-hydroxylase or non-heme iron oxygenase | ||
| + | |||
| + | **[[4m23]] – SlORFP – ''Streptomyces lavendulae''<br /> | ||
| + | **[[4m2i]] – SlORFP + Fe <br /> | ||
| + | **[[4m25]] – SlORFP + Fe + ketoglutarate <br /> | ||
| + | **[[4m27]], [[4m2c]] – SlORFP + Fe + Arg<br /> | ||
| + | **[[4m2e]] – SlORFP + Fe + Arg derivative<br /> | ||
| + | **[[4m2f]] – SlORFP + Fe + canavanine<br /> | ||
| + | **[[4m2g]], [[4m26]] – SlORFP + Fe + succinate + Arg derivative<br /> | ||
| + | **[[2wbo]] – SvVIOC + Fe + Arg – ''Streptomyces vinaceus''<br /> | ||
| + | **[[2wbq]] – SvVIOC + Arg derivative<br /> | ||
| + | **[[2wbp]] – SvVIOC + Fe + succinate + Arg derivative<br /> | ||
*Prolyl 4-hydroxylase | *Prolyl 4-hydroxylase | ||
| + | **[[4bt8]], [[2yq8]] - hPH a1 subunit collagen-binding domain<br /> | ||
| + | **[[1tjc]] - hPH a1 subunit collagen-binding domain (mutant)<br /> | ||
| + | **[[4btb]], [[4bta]], [[4bt9]] - hPH a1 subunit + peptide<br /> | ||
| + | **[[6evl]] - hPH a2 subunit <br /> | ||
| + | **[[6evm]], [[6evn]], [[6evo]], [[6evp]] - hPH a2 subunit + peptide<br /> | ||
**[[5hv0]] - PH + Cd – ''Bacillus anthracis'' <br /> | **[[5hv0]] - PH + Cd – ''Bacillus anthracis'' <br /> | ||
**[[5c5u]] - PcvPH + Mn – Paramecium chlorella virus <br /> | **[[5c5u]] - PcvPH + Mn – Paramecium chlorella virus <br /> | ||
| Line 139: | Line 172: | ||
**[[4p7x]] - RlPH + Co + pipecolic acid – ''Rhizobium loti'' <br /> | **[[4p7x]] - RlPH + Co + pipecolic acid – ''Rhizobium loti'' <br /> | ||
**[[4p7w]] - RlPH + Co + proline<br /> | **[[4p7w]] - RlPH + Co + proline<br /> | ||
| + | **[[6iuq]] - PH + Fe – ''Phytophthora capsici''<br /> | ||
| + | **[[6ey1]] - TaPH + Mn – ''Trichoplax adhaerens''<br /> | ||
| + | **[[6f0w]] - TaPH + Mn + hypoxia inducible factor + oxalylglycine <br /> | ||
| + | **[[2v4a]], [[2jij]] - CrPH – ''Chlamydomonas reinhardtii'' <br /> | ||
| + | **[[2jig]] - CrPH + Zn + pyridine derivative<br /> | ||
| + | |||
| + | *HIF prolyl hydroxylase 1 or egl nine homolog 2 | ||
| + | |||
| + | **[[5v1b]] – hHPH-1 + Fe + inhibitor <br /> | ||
| + | |||
| + | *HIF prolyl hydroxylase 2 or egl nine homolog 1 | ||
| + | |||
| + | **[[5las]] – hHPH-2 + HIF <br /> | ||
| + | **[[5l9b]] – hHPH-2 catalytic domain (mutant) + Mn + HIF + oxoglutarate <br /> | ||
| + | **[[5l9v]], [[5la9]], [[3hqr]], [[3hqu]] – hHPH-2 catalytic domain (mutant) + Mn + HIF + oxalylglycine <br /> | ||
| + | **[[5ox5]], [[5ox6]], [[4bqw]], [[4bqx]], [[4bqy]] – hHPH-2 catalytic domain + Mn + inhibitor <br /> | ||
| + | **[[5l9r]] – hHPH-2 catalytic domain + Mn + oxalylglycine <br /> | ||
| + | **[[5v18]], [[6nmq]], [[3ouh]], [[3oui]], [[3ouj]], [[2y34]], [[2hbt]], [[2hbu]], [[2g19]], [[2g1m]] – hHPH-2 catalytic domain + Fe + inhibitor <br /> | ||
| + | **[[4jzr]] – hHPH-2 catalytic domain + Ni + inhibitor <br /> | ||
| + | **[[2y33]] – hHPH-2 catalytic domain + Zn + inhibitor <br /> | ||
| + | **[[5lat]], [[5lb6]], [[5lbb]], [[5lbc]], [[5lbe]], [[5lbf]], [[5a3u]], [[4uwd]], [[4kbz]] – hHPH-2 catalytic domain (mutant) + Mn + inhibitor <br /> | ||
| + | |||
| + | *Lysine 4-hydroxylase | ||
| + | |||
| + | **[[6f9p]] – FlL4H + Fe – ''Flavobacterium'' <br /> | ||
| + | **[[6exf]] – FlL4H + Fe + lysine <br /> | ||
| + | **[[6euo]] – FlL4H + Fe + succinate derivative<br /> | ||
| + | **[[6eur]] – FlL4H + Fe + oxoglutarate<br /> | ||
| + | **[[6exh]] – FlL4H + Fe + lysine + succinate<br /> | ||
| + | |||
| + | *Lysine 3-hydroxylase | ||
| + | |||
| + | **[[6f2e]] – CaL3H – ''Catenulispora acidiphila'' <br /> | ||
| + | **[[6f2b]] – CaL3H + Fe + oxoglutarate <br /> | ||
| + | **[[6f2a]] – CaL3H + Fe + lysine <br /> | ||
| + | **[[6f6j]] – CaL3H + Fe + succinate + hydroxyl lysine <br /> | ||
*Salicilate hydroxylase | *Salicilate hydroxylase | ||
Revision as of 09:43, 25 August 2019
| |||||||||||
3D structures of hydroxylases
Updated on 25-August-2019
Additional Resources
For additional information, see: Amino Acid Synthesis & Metabolism
References
- ↑ Fitzpatrick PF. The aromatic amino acid hydroxylases. Adv Enzymol Relat Areas Mol Biol. 2000;74:235-94. PMID:10800597
- ↑ KAUFMAN S. The enzymatic conversion of phenylalanine to tyrosine. J Biol Chem. 1957 May;226(1):511-24. PMID:13428782
- ↑ Chen D, Frey PA. Phenylalanine hydroxylase from Chromobacterium violaceum. Uncoupled oxidation of tetrahydropterin and the role of iron in hyroxylation. J Biol Chem. 1998 Oct 2;273(40):25594-601. PMID:9748224
- ↑ Leiros HK, Pey AL, Innselset M, Moe E, Leiros I, Steen IH, Martinez A. Structure of phenylalanine hydroxylase from Colwellia psychrerythraea 34H, a monomeric cold active enzyme with local flexibility around the active site and high overall stability. J Biol Chem. 2007 Jul 27;282(30):21973-86. Epub 2007 May 30. PMID:17537732 doi:10.1074/jbc.M610174200
- ↑ Grenett HE, Ledley FD, Reed LL, Woo SL. Full-length cDNA for rabbit tryptophan hydroxylase: functional domains and evolution of aromatic amino acid hydroxylases. Proc Natl Acad Sci U S A. 1987 Aug;84(16):5530-4. PMID:3475690
- ↑ NAGATSU T, LEVITT M, UDENFRIEND S. TYROSINE HYDROXYLASE. THE INITIAL STEP IN NOREPINEPHRINE BIOSYNTHESIS. J Biol Chem. 1964 Sep;239:2910-7. PMID:14216443
- ↑ 7.0 7.1 LEVINE RJ, LOVENBERG W, SJOERDSMA A. HYDROXYLATION OF TRYPTOPHAN AND PHENYLALANINE IN NEOPLASTIC MAST CELLS OF THE MOUSE. Biochem Pharmacol. 1964 Sep;13:1283-90. PMID:14221726
- ↑ Grahame-Smith DG. Tryptophan hydroxylation in brain. Biochem Biophys Res Commun. 1964 Aug 11;16(6):586-92. PMID:5297063
- ↑ 9.0 9.1 Walther DJ, Bader M. A unique central tryptophan hydroxylase isoform. Biochem Pharmacol. 2003 Nov 1;66(9):1673-80. PMID:14563478
- ↑ Grenett HE, Ledley FD, Reed LL, Woo SL. Full-length cDNA for rabbit tryptophan hydroxylase: functional domains and evolution of aromatic amino acid hydroxylases. Proc Natl Acad Sci U S A. 1987 Aug;84(16):5530-4. PMID:3475690
- ↑ Craig SP, Boularand S, Darmon MC, Mallet J, Craig IW. Localization of human tryptophan hydroxylase (TPH) to chromosome 11p15.3----p14 by in situ hybridization. Cytogenet Cell Genet. 1991;56(3-4):157-9. PMID:2055111
- ↑ Walther DJ, Peter JU, Bashammakh S, Hortnagl H, Voits M, Fink H, Bader M. Synthesis of serotonin by a second tryptophan hydroxylase isoform. Science. 2003 Jan 3;299(5603):76. PMID:12511643 doi:http://dx.doi.org/10.1126/science.1078197
- ↑ 13.0 13.1 Patel PD, Pontrello C, Burke S. Robust and tissue-specific expression of TPH2 versus TPH1 in rat raphe and pineal gland. Biol Psychiatry. 2004 Feb 15;55(4):428-33. PMID:14960297 doi:http://dx.doi.org/10.1016/j.biopsych.2003.09.002
- ↑ Slominski A, Pisarchik A, Johansson O, Jing C, Semak I, Slugocki G, Wortsman J. Tryptophan hydroxylase expression in human skin cells. Biochim Biophys Acta. 2003 Oct 15;1639(2):80-6. PMID:14559114
- ↑ Hasegawa H, Yanagisawa M, Inoue F, Yanaihara N, Ichiyama A. Demonstration of non-neural tryptophan 5-mono-oxygenase in mouse intestinal mucosa. Biochem J. 1987 Dec 1;248(2):501-9. PMID:3435461
- ↑ Hosoda S, Nakamura W, Takatsuki K. Properties of tryptophan hydroxylase from human carcinoid tumor. Biochim Biophys Acta. 1977 May 12;482(1):27-34. PMID:16654
- ↑ Manuck SB, Flory JD, Ferrell RE, Dent KM, Mann JJ, Muldoon MF. Aggression and anger-related traits associated with a polymorphism of the tryptophan hydroxylase gene. Biol Psychiatry. 1999 Mar 1;45(5):603-14. PMID:10088047
- ↑ Mitchell JJ, Trakadis YJ, Scriver CR. Phenylalanine hydroxylase deficiency. Genet Med. 2011 Aug;13(8):697-707. doi: 10.1097/GIM.0b013e3182141b48. PMID:21555948 doi:http://dx.doi.org/10.1097/GIM.0b013e3182141b48
- ↑ Flatmark T, Stevens RC. Structural Insight into the Aromatic Amino Acid Hydroxylases and Their Disease-Related Mutant Forms. Chem Rev. 1999 Aug 11;99(8):2137-2160. PMID:11849022
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, Michael Skovbo Windahl, David Canner, Joel L. Sussman, Jaime Prilusky