This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Hemeproteins
From Proteopedia
(Difference between revisions)
| Line 120: | Line 120: | ||
The <scene name='46/466465/Cv/3'>O2 molecule in the active site coordinates with the heme</scene> (PDB entry [[1dcc]]).<ref>PMID:7664080</ref> Water molecules are shown as red spheres. | The <scene name='46/466465/Cv/3'>O2 molecule in the active site coordinates with the heme</scene> (PDB entry [[1dcc]]).<ref>PMID:7664080</ref> Water molecules are shown as red spheres. | ||
| - | =Cytochrome P450 hydroxylase= | ||
| - | '''Cytochrome P450 hydroxylase''' (CPH) acts in the in-chain hydroxylation of lauric acid which is required for the development of the male organ in higher plants<ref>PMID:24798002</ref>. CPH hydroxylyzes the anti-cholesterol natural product herbosidiene<ref>PMID:25139567</ref>. | ||
| - | *<scene name='77/774654/Cv/6'>Lauric acid binding site</scene>. | ||
| - | *<scene name='77/774654/Cv/7'>Heme group binding site</scene>. | ||
| - | *<scene name='77/774654/Cv/8'>Fe coordination site</scene>. | ||
| - | *<scene name='77/774654/Cv/9'>Distances between Lauric acid oxygens and heme group Fe</scene> in Cytochrome P450 hydroxylase from ''Steptomyces avermitilis'' (PDB code [[5cwe]]). | ||
=Cytochrome c Nitrite Reductase= | =Cytochrome c Nitrite Reductase= | ||
| Line 136: | Line 130: | ||
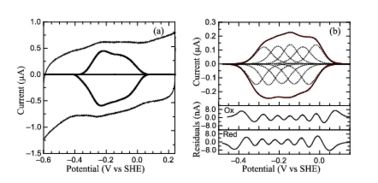
[[Image:figur5.jpg|left|378px|thumb|PFV of ''S. oneidensis'' ccNiR (a) Typical signal on a graphite electrode. (b) Baselinesubtracted non-turnover voltammogram]] | [[Image:figur5.jpg|left|378px|thumb|PFV of ''S. oneidensis'' ccNiR (a) Typical signal on a graphite electrode. (b) Baselinesubtracted non-turnover voltammogram]] | ||
The Ca<sup>2+</sup> ion within <scene name='Journal:JBIC:16/Cv/14'>conserved site</scene> is coordinated in bidentate fashion by <scene name='Journal:JBIC:16/Cv/15'>Glu205</scene>, and in monodentate fashion by the <scene name='Journal:JBIC:16/Cv/16'>Tyr206 and Lys254</scene> backbone carbonyls, and the <scene name='Journal:JBIC:16/Cv/17'>Gln256</scene> side-chain carbonyl. In the ''S. oneidensis'' structure only <scene name='Journal:JBIC:16/Cv/18'>one water molecule</scene> is assigned to the Ca<sup>2+</sup> ion in subunit B. In subunit A the difference electron density that represents this water molecule is very close to the noise level, and it is difficult to identify even one water molecule there. The <scene name='Journal:JBIC:16/Cv/14'>carbonyl side chain of Asp242 and the hydroxyl of Tyr235</scene> come near to the open calcium coordination sites, but are not within bonding distance. Instead they interact with the water molecule that is weakly coordinated to the Ca<sup>2+</sup> ion. The ccNiR calcium ions appear to play a vital role in organizing the <scene name='Journal:JBIC:16/Cv/13'>active site</scene> (as was mentioned above <font color='magenta'><b>hemes-1</b></font> are the active sites). | The Ca<sup>2+</sup> ion within <scene name='Journal:JBIC:16/Cv/14'>conserved site</scene> is coordinated in bidentate fashion by <scene name='Journal:JBIC:16/Cv/15'>Glu205</scene>, and in monodentate fashion by the <scene name='Journal:JBIC:16/Cv/16'>Tyr206 and Lys254</scene> backbone carbonyls, and the <scene name='Journal:JBIC:16/Cv/17'>Gln256</scene> side-chain carbonyl. In the ''S. oneidensis'' structure only <scene name='Journal:JBIC:16/Cv/18'>one water molecule</scene> is assigned to the Ca<sup>2+</sup> ion in subunit B. In subunit A the difference electron density that represents this water molecule is very close to the noise level, and it is difficult to identify even one water molecule there. The <scene name='Journal:JBIC:16/Cv/14'>carbonyl side chain of Asp242 and the hydroxyl of Tyr235</scene> come near to the open calcium coordination sites, but are not within bonding distance. Instead they interact with the water molecule that is weakly coordinated to the Ca<sup>2+</sup> ion. The ccNiR calcium ions appear to play a vital role in organizing the <scene name='Journal:JBIC:16/Cv/13'>active site</scene> (as was mentioned above <font color='magenta'><b>hemes-1</b></font> are the active sites). | ||
| + | |||
| + | =Cytochrome P450 hydroxylase= | ||
| + | '''Cytochrome P450 hydroxylase''' (CPH) acts in the in-chain hydroxylation of lauric acid which is required for the development of the male organ in higher plants<ref>PMID:24798002</ref>. CPH hydroxylyzes the anti-cholesterol natural product herbosidiene<ref>PMID:25139567</ref>. | ||
| + | *<scene name='77/774654/Cv/6'>Lauric acid binding site</scene>. | ||
| + | *<scene name='77/774654/Cv/7'>Heme group binding site</scene>. | ||
| + | *<scene name='77/774654/Cv/8'>Fe coordination site</scene>. | ||
| + | *<scene name='77/774654/Cv/9'>Distances between Lauric acid oxygens and heme group Fe</scene> in Cytochrome P450 hydroxylase from ''Steptomyces avermitilis'' (PDB code [[5cwe]]). | ||
=[[Hemoglobin]]= | =[[Hemoglobin]]= | ||
=[[Myoglobin]]= | =[[Myoglobin]]= | ||
Revision as of 15:20, 11 November 2019
| |||||||||||
References
- ↑ Schenkman JB, Jansson I. The many roles of cytochrome b5. Pharmacol Ther. 2003 Feb;97(2):139-52. PMID:12559387
- ↑ Rodriguez-Maranon MJ, Qiu F, Stark RE, White SP, Zhang X, Foundling SI, Rodriguez V, Schilling CL 3rd, Bunce RA, Rivera M. 13C NMR spectroscopic and X-ray crystallographic study of the role played by mitochondrial cytochrome b5 heme propionates in the electrostatic binding to cytochrome c. Biochemistry. 1996 Dec 17;35(50):16378-90. PMID:8973214 doi:10.1021/bi961895o
- ↑ Crofts AR. The cytochrome bc1 complex: function in the context of structure. Annu Rev Physiol. 2004;66:689-733. PMID:14977419 doi:http://dx.doi.org/10.1146/annurev.physiol.66.032102.150251
- ↑ Berry EA, Huang LS, Saechao LK, Pon NG, Valkova-Valchanova M, Daldal F. X-Ray Structure of Rhodobacter Capsulatus Cytochrome bc (1): Comparison with its Mitochondrial and Chloroplast Counterparts. Photosynth Res. 2004;81(3):251-75. PMID:16034531 doi:http://dx.doi.org/10.1023/B:PRES.0000036888.18223.0e
- ↑ Rajagopal BS, Wilson MT, Bendall DS, Howe CJ, Worrall JA. Structural and kinetic studies of imidazole binding to two members of the cytochrome c (6) family reveal an important role for a conserved heme pocket residue. J Biol Inorg Chem. 2011 Jan 26. PMID:21267610 doi:10.1007/s00775-011-0758-y
- ↑ Morelli X, Czjzek M, Hatchikian CE, Bornet O, Fontecilla-Camps JC, Palma NP, Moura JJ, Guerlesquin F. Structural model of the Fe-hydrogenase/cytochrome c553 complex combining transverse relaxation-optimized spectroscopy experiments and soft docking calculations. J Biol Chem. 2000 Jul 28;275(30):23204-10. PMID:10748163 doi:10.1074/jbc.M909835199
- ↑ Manole A, Kekilli D, Svistunenko DA, Wilson MT, Dobbin PS, Hough MA. Conformational control of the binding of diatomic gases to cytochrome c'. J Biol Inorg Chem. 2015 Mar 20. PMID:25792378 doi:http://dx.doi.org/10.1007/s00775-015-1253-7
- ↑ 8.00 8.01 8.02 8.03 8.04 8.05 8.06 8.07 8.08 8.09 8.10 8.11 8.12 Stelter M, Melo AM, Pereira MM, Gomes CM, Hreggvidsson GO, Hjorleifsdottir S, Saraiva LM, Teixeira M, Archer M. A Novel Type of Monoheme Cytochrome c: Biochemical and Structural Characterization at 1.23 A Resolution of Rhodothermus marinus Cytochrome c. Biochemistry. 2008 Oct 15. PMID:18855424 doi:10.1021/bi800999g
- ↑ Than ME, Hof P, Huber R, Bourenkov GP, Bartunik HD, Buse G, Soulimane T. Thermus thermophilus cytochrome-c552: A new highly thermostable cytochrome-c structure obtained by MAD phasing. J Mol Biol. 1997 Aug 29;271(4):629-44. PMID:9281430 doi:10.1006/jmbi.1997.1181
- ↑ Soares CM, Baptista AM, Pereira MM, Teixeira M. Investigation of protonatable residues in Rhodothermus marinus caa3 haem-copper oxygen reductase: comparison with Paracoccus denitrificans aa3 haem-copper oxygen reductase. J Biol Inorg Chem. 2004 Mar;9(2):124-34. Epub 2003 Dec 23. PMID:14691678 doi:10.1007/s00775-003-0509-9
- ↑ Pereira MM, Santana M, Teixeira M. A novel scenario for the evolution of haem-copper oxygen reductases. Biochim Biophys Acta. 2001 Jun 1;1505(2-3):185-208. PMID:11334784
- ↑ 12.0 12.1 12.2 12.3 12.4 12.5 Karp, Gerald (2008). Cell and Molecular Biology (5th edition). Hoboken, NJ: John Wiley & Sons. ISBN 978-0470042175.
- ↑ Prince RC, George GN. Cytochrome f revealed. Trends Biochem Sci. 1995 Jun;20(6):217-8. PMID:7631417
- ↑ Martinez SE, Huang D, Szczepaniak A, Cramer WA, Smith JL. Crystal structure of chloroplast cytochrome f reveals a novel cytochrome fold and unexpected heme ligation. Structure. 1994 Feb 15;2(2):95-105. PMID:8081747
- ↑ Danielson PB. The cytochrome P450 superfamily: biochemistry, evolution and drug metabolism in humans. Curr Drug Metab. 2002 Dec;3(6):561-97. PMID:12369887
- ↑ Williams PA, Cosme J, Ward A, Angove HC, Matak Vinkovic D, Jhoti H. Crystal structure of human cytochrome P450 2C9 with bound warfarin. Nature. 2003 Jul 24;424(6947):464-8. Epub 2003 Jul 13. PMID:12861225 doi:http://dx.doi.org/10.1038/nature01862
- ↑ Tsukihara T, Aoyama H, Yamashita E, Tomizaki T, Yamaguchi H, Shinzawa-Itoh K, Nakashima R, Yaono R, Yoshikawa S. Structures of metal sites of oxidized bovine heart cytochrome c oxidase at 2.8 A. Science. 1995 Aug 25;269(5227):1069-74. PMID:7652554
- ↑ Yoshikawa S, Shinzawa-Itoh K, Nakashima R, Yaono R, Yamashita E, Inoue N, Yao M, Fei MJ, Libeu CP, Mizushima T, Yamaguchi H, Tomizaki T, Tsukihara T. Redox-coupled crystal structural changes in bovine heart cytochrome c oxidase. Science. 1998 Jun 12;280(5370):1723-9. PMID:9624044
- ↑ Atack JM, Kelly DJ. Structure, mechanism and physiological roles of bacterial cytochrome c peroxidases. Adv Microb Physiol. 2007;52:73-106. PMID:17027371 doi:http://dx.doi.org/10.1016/S0065-2911(06)52002-8
- ↑ Miller MA, Shaw A, Kraut J. 2.2 A structure of oxy-peroxidase as a model for the transient enzyme: peroxide complex. Nat Struct Biol. 1994 Aug;1(8):524-31. PMID:7664080
- ↑ 21.0 21.1 none yet
- ↑ Yang X, Wu D, Shi J, He Y, Pinot F, Grausem B, Yin C, Zhu L, Chen M, Luo Z, Liang W, Zhang D. Rice CYP703A3, a cytochrome P450 hydroxylase, is essential for development of anther cuticle and pollen exine. J Integr Plant Biol. 2014 Oct;56(10):979-94. doi: 10.1111/jipb.12212. Epub 2014, Jul 15. PMID:24798002 doi:http://dx.doi.org/10.1111/jipb.12212
- ↑ Yu D, Xu F, Shao L, Zhan J. A specific cytochrome P450 hydroxylase in herboxidiene biosynthesis. Bioorg Med Chem Lett. 2014 Sep 15;24(18):4511-4514. doi:, 10.1016/j.bmcl.2014.07.078. Epub 2014 Aug 6. PMID:25139567 doi:http://dx.doi.org/10.1016/j.bmcl.2014.07.078