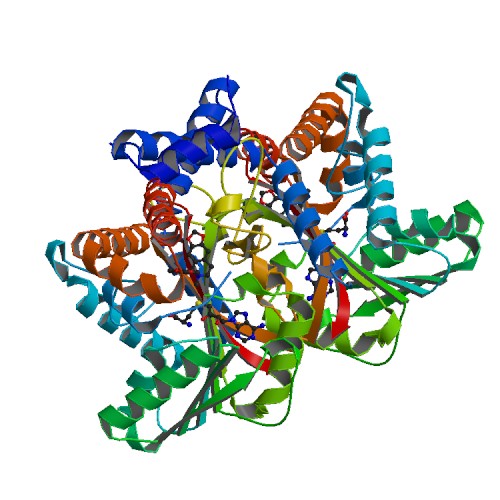
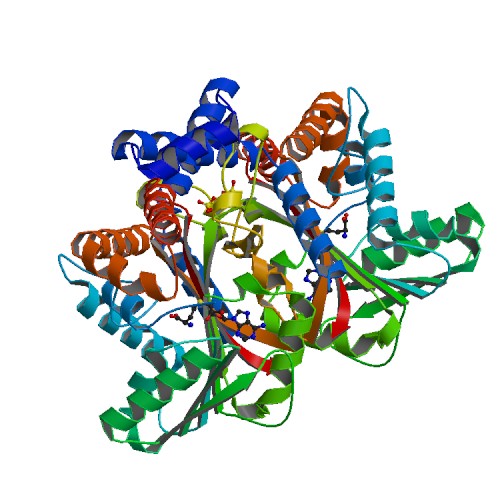
Tetrahydroprotoberberine N-methyltransferase
From Proteopedia
(Difference between revisions)
| Line 13: | Line 13: | ||
This protein has a <scene name='82/829888/Catalytic_triad/7'>catalytic triad</scene> which consists of amino acids His-208, Glu-204, and Glu-207. The authors explained within the paper that other amino acids may play a role in the triad as well. They were unsure but those three were the most accurate. These amino acids play an important role in catalysis for the protein. <ref> Bennett, M. R., Thompson, M. L., Shepherd, S. A., Dunstan, M. S., Herbert, A. J., Smith, D. R. M., Cronin, V. A., Menon, B. R. K., Levy, C., and Micklefield, J. (2018) Structure and biocatalytic scope of coclaurine Nmethyltransferase.Angew. Chem. Int. Ed. Engl. 57, 10600–10604CrossRef | This protein has a <scene name='82/829888/Catalytic_triad/7'>catalytic triad</scene> which consists of amino acids His-208, Glu-204, and Glu-207. The authors explained within the paper that other amino acids may play a role in the triad as well. They were unsure but those three were the most accurate. These amino acids play an important role in catalysis for the protein. <ref> Bennett, M. R., Thompson, M. L., Shepherd, S. A., Dunstan, M. S., Herbert, A. J., Smith, D. R. M., Cronin, V. A., Menon, B. R. K., Levy, C., and Micklefield, J. (2018) Structure and biocatalytic scope of coclaurine Nmethyltransferase.Angew. Chem. Int. Ed. Engl. 57, 10600–10604CrossRef | ||
Medline </ref>. The basic <scene name='82/829888/Spacefill_view_of_protein/1'>spacefill view</scene> of the entire protein alllows readers to visualize the different elements show in different colors. The elements shown are carbons(grey), nitrogen(blue), and oxygen(red). This protein has a <scene name='82/829888/Ligand/1'>ligand</scene> which is SAM. There are <scene name='82/829888/Hydrophilic_side_chains/1'>hydrophilic side chains</scene>of SAM that form a small catalytic pocket and surrounds the amino group and methyl donor of SAM. This catalytic pocket forms a L shape. In green Glu-204, yellow is Glu-207, red is His-208, and Tyr-81 is blue. The <scene name='82/829888/Active_site/3'>active site</scene> of the protein consists of amino acids Valine-188(yellow), Aspartic Acid-187(blue), and Alanine-186(green), with purple being the rest of the ligand, SAM. The active site is the region where substrate molecules bind and undergo a chemical reaction. The <scene name='82/829888/Secondary_structure/1'>secondary structure</scene> of this protein contains a pattern of hydrogen bonds between atoms in the peptide bond. This cartoon view allows readers to visualize the alpha(pink) and beta sheets(yellow). The protein consists of two regions which are <scene name='82/829888/Polar_and_non-polar/1'>polar and non-polar regions</scene>. The non-polar region is in grey while the polar region is in purple. The <scene name='82/829888/Cationic_region/1'>cationic region</scene> of the protein has a side chain of Lysine, Arginine, Aspartic Acid, and Glutamic Acid. The cationic region is in the light blue. | Medline </ref>. The basic <scene name='82/829888/Spacefill_view_of_protein/1'>spacefill view</scene> of the entire protein alllows readers to visualize the different elements show in different colors. The elements shown are carbons(grey), nitrogen(blue), and oxygen(red). This protein has a <scene name='82/829888/Ligand/1'>ligand</scene> which is SAM. There are <scene name='82/829888/Hydrophilic_side_chains/1'>hydrophilic side chains</scene>of SAM that form a small catalytic pocket and surrounds the amino group and methyl donor of SAM. This catalytic pocket forms a L shape. In green Glu-204, yellow is Glu-207, red is His-208, and Tyr-81 is blue. The <scene name='82/829888/Active_site/3'>active site</scene> of the protein consists of amino acids Valine-188(yellow), Aspartic Acid-187(blue), and Alanine-186(green), with purple being the rest of the ligand, SAM. The active site is the region where substrate molecules bind and undergo a chemical reaction. The <scene name='82/829888/Secondary_structure/1'>secondary structure</scene> of this protein contains a pattern of hydrogen bonds between atoms in the peptide bond. This cartoon view allows readers to visualize the alpha(pink) and beta sheets(yellow). The protein consists of two regions which are <scene name='82/829888/Polar_and_non-polar/1'>polar and non-polar regions</scene>. The non-polar region is in grey while the polar region is in purple. The <scene name='82/829888/Cationic_region/1'>cationic region</scene> of the protein has a side chain of Lysine, Arginine, Aspartic Acid, and Glutamic Acid. The cationic region is in the light blue. | ||
| + | |||
| + | = Energy Transformation = | ||
| + | There isn't any energy transformation data presented in the paper. | ||
</StructureSection> | </StructureSection> | ||
== References == | == References == | ||
<references/> | <references/> | ||
Revision as of 04:43, 1 December 2019
Structure
| |||||||||||
References
- ↑ Takao, N., Kamigauchi, M., and Okada, M. (1983) Biosynthesis of benzo-[c]phenanthridine alkaloids sanguinarine, chelirubine and macarpine.Helv. Chim. Acta 66, 473–484 CrossRef
- ↑ Bennett, M. R., Thompson, M. L., Shepherd, S. A., Dunstan, M. S., Herbert, A. J., Smith, D. R. M., Cronin, V. A., Menon, B. R. K., Levy, C., and Micklefield, J. (2018) Structure and biocatalytic scope of coclaurine Nmethyltransferase.Angew. Chem. Int. Ed. Engl. 57, 10600–10604CrossRef Medline