This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Harrison L. Smith/Sandbox1
From Proteopedia
(Difference between revisions)
| Line 21: | Line 21: | ||
The insulin receptor is a receptor tyrosine kinase that exists in two stable conformations, an inactive and active state. The entire insulin receptor is a dimer of heterodimers with two extracellular alpha subunits, and two transmembrane/intracellular beta subunits linked and stabilized by multiple disulfide bonds. | The insulin receptor is a receptor tyrosine kinase that exists in two stable conformations, an inactive and active state. The entire insulin receptor is a dimer of heterodimers with two extracellular alpha subunits, and two transmembrane/intracellular beta subunits linked and stabilized by multiple disulfide bonds. | ||
======Alpha Subunit====== | ======Alpha Subunit====== | ||
| - | The alpha subunit is the extracellular domain of the insulin receptor and is the site of insulin binding. The alpha subunit is comprised of two Leucine rich domains (L1 & L2), a | + | The alpha subunit is the extracellular domain of the insulin receptor and is the site of insulin binding. The alpha subunit is comprised of two Leucine rich domains (L1 & L2), a Cysteine rich domain (CR), and a C-terminal alpha helix. The actual site of insulin binding occurs at the C-terminal alpha helix and is stabilized by the L1 and L2 domains. |
======Beta Subunit====== | ======Beta Subunit====== | ||
| - | The beta subunit is the membrane spanning and intracellular portion of the insulin receptor. This domain is composed of | + | The beta subunit is the membrane spanning and intracellular portion of the insulin receptor. This domain is composed of three Fibronectin domains (FN III-1,-2,-3) and the tyrosine kinase domain. The tyrosine kinase domain is the site of autophosphorylation upon activation of the receptor. |
===Conformation Change=== | ===Conformation Change=== | ||
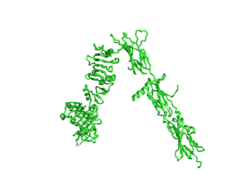
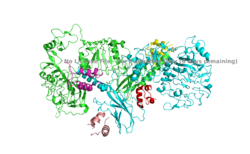
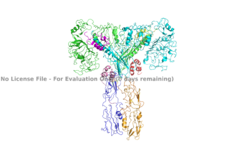
The inactive form of the insulin receptor predominates in low-levels of circulating insulin, whereas the active conformation is seen when insulin binds to any of the 4 receptor sites. The inactive conformation resembles an inverted V (Figure 1), and the active conformation resembles a T (Figure 2). Upon the binding of insulin to any of the four binding sites, the conformation change will begin, causing the Beta subunit's tyrosine kinase domains to move close together, allowing them to autophosphorylate. This autophosphorylation is what activates the insulin receptor and allows it to participate in further downstream signaling pathways. <ref> DOI 10.1038/s41467-018-06826-6</ref>. | The inactive form of the insulin receptor predominates in low-levels of circulating insulin, whereas the active conformation is seen when insulin binds to any of the 4 receptor sites. The inactive conformation resembles an inverted V (Figure 1), and the active conformation resembles a T (Figure 2). Upon the binding of insulin to any of the four binding sites, the conformation change will begin, causing the Beta subunit's tyrosine kinase domains to move close together, allowing them to autophosphorylate. This autophosphorylation is what activates the insulin receptor and allows it to participate in further downstream signaling pathways. <ref> DOI 10.1038/s41467-018-06826-6</ref>. | ||
Revision as of 21:43, 29 March 2020
Homo sapiens Insulin Receptor Ectodomain
| |||||||||||
References
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644
- ↑ Tatulian SA. Structural Dynamics of Insulin Receptor and Transmembrane Signaling. Biochemistry. 2015 Sep 15;54(36):5523-32. doi: 10.1021/acs.biochem.5b00805. Epub , 2015 Sep 3. PMID:26322622 doi:http://dx.doi.org/10.1021/acs.biochem.5b00805
- ↑ Weis F, Menting JG, Margetts MB, Chan SJ, Xu Y, Tennagels N, Wohlfart P, Langer T, Muller CW, Dreyer MK, Lawrence MC. The signalling conformation of the insulin receptor ectodomain. Nat Commun. 2018 Oct 24;9(1):4420. doi: 10.1038/s41467-018-06826-6. PMID:30356040 doi:http://dx.doi.org/10.1038/s41467-018-06826-6
- ↑ Uchikawa E, Choi E, Shang G, Yu H, Bai XC. Activation mechanism of the insulin receptor revealed by cryo-EM structure of the fully liganded receptor-ligand complex. Elife. 2019 Aug 22;8. pii: 48630. doi: 10.7554/eLife.48630. PMID:31436533 doi:http://dx.doi.org/10.7554/eLife.48630
- ↑ Wilcox G. Insulin and insulin resistance. Clin Biochem Rev. 2005 May;26(2):19-39. PMID:16278749
- ↑ Riddle MC. Treatment of diabetes with insulin. From art to science. West J Med. 1983 Jun;138(6):838-46. PMID:6351440
Student Contributors
- Harrison Smith
- Alyssa Ritter