Sandbox Reserved 1600
From Proteopedia
(Difference between revisions)
| Line 6: | Line 6: | ||
=Introduction= | =Introduction= | ||
| - | Bd oxidase is an integral membrane protein that catalyzes the reduction of | + | Bd oxidase is an integral membrane protein that catalyzes the reduction of O₂ -> 2H₂O using quinol as the reducing substrate. <ref name=”Giuffrè”>PMID:24486503</ref> The reaction is electrogenic but is not coupled to a proton pump it instead uses internal water molecules to provide the protons needed for the reduction reaction. <ref name=”Safarian”>PMID:31604309</ref> It plays a key role in protecting the organism from high oxidative stress (ie. preventing free radicals in intracellular space in prokaryotes, more specifically gram-negative heterotrophs). <ref name=”Jünemann”>PMID:9332500</ref> |
There are two main types of respiratory cytochrome oxidases: the heme/copper oxidases, and the heme-only cytochrome bd quinol oxidase, which is what bd oxidase falls under. <ref name=”Das”>PMID:15743950</ref> Heme-only cytochrome bd quinol oxidases are associated with microaerobic dioxygen respiration, and they have a high affinity for oxygen. | There are two main types of respiratory cytochrome oxidases: the heme/copper oxidases, and the heme-only cytochrome bd quinol oxidase, which is what bd oxidase falls under. <ref name=”Das”>PMID:15743950</ref> Heme-only cytochrome bd quinol oxidases are associated with microaerobic dioxygen respiration, and they have a high affinity for oxygen. | ||
| Line 34: | Line 34: | ||
Another potential entry site is related to the a1-4 four-helix bundle of CydB. Therefore, this is called the CydB pathway. In this pathway, Asp25 is thought to be the equivalent of the Glu108 in the CydA pathway <ref name=”Safarian”>PMID: 27126043 </ref>. The other residues help facilitate the movement of the proton very similarly to the CydA pathway. There is less known about the CydB pathway, and therefore, the CydA pathway is the most accepted source of protons. | Another potential entry site is related to the a1-4 four-helix bundle of CydB. Therefore, this is called the CydB pathway. In this pathway, Asp25 is thought to be the equivalent of the Glu108 in the CydA pathway <ref name=”Safarian”>PMID: 27126043 </ref>. The other residues help facilitate the movement of the proton very similarly to the CydA pathway. There is less known about the CydB pathway, and therefore, the CydA pathway is the most accepted source of protons. | ||
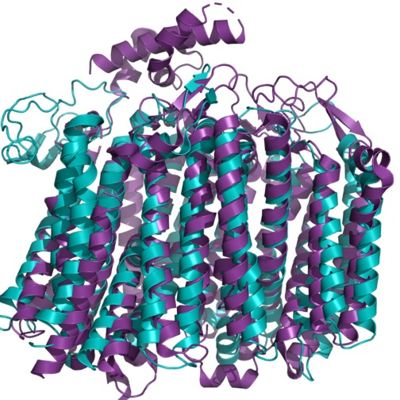
| - | == | + | == Structure Similarity to Bd Oxidase found in ''Ecoli'' == |
| + | [[Image:Aligmentbdoidase.jpg|400 px|right|thumb|Figure 1. Alignment of bd oxidase for the organisms ''Geobacillus thermodenitrificans'' (blue) and ''Ecoli'' (purple).]] | ||
| - | == Structural highlights == | ||
| - | |||
| - | This is a sample scene created with SAT to <scene name="/12/3456/Sample/1">color</scene> by Group, and another to make <scene name="/12/3456/Sample/2">a transparent representation</scene> of the protein. You can make your own scenes on SAT starting from scratch or loading and editing one of these sample scenes. | ||
</StructureSection> | </StructureSection> | ||
== References == | == References == | ||
<references/> | <references/> | ||
Revision as of 21:56, 29 March 2020
bd oxidase; Geobacillus thermodenitrificans
| |||||||||||
References
- ↑ Giuffre A, Borisov VB, Arese M, Sarti P, Forte E. Cytochrome bd oxidase and bacterial tolerance to oxidative and nitrosative stress. Biochim Biophys Acta. 2014 Jul;1837(7):1178-87. doi:, 10.1016/j.bbabio.2014.01.016. Epub 2014 Jan 31. PMID:24486503 doi:http://dx.doi.org/10.1016/j.bbabio.2014.01.016
- ↑ Safarian S, Hahn A, Mills DJ, Radloff M, Eisinger ML, Nikolaev A, Meier-Credo J, Melin F, Miyoshi H, Gennis RB, Sakamoto J, Langer JD, Hellwig P, Kuhlbrandt W, Michel H. Active site rearrangement and structural divergence in prokaryotic respiratory oxidases. Science. 2019 Oct 4;366(6461):100-104. doi: 10.1126/science.aay0967. PMID:31604309 doi:http://dx.doi.org/10.1126/science.aay0967
- ↑ Junemann S. Cytochrome bd terminal oxidase. Biochim Biophys Acta. 1997 Aug 22;1321(2):107-27. doi:, 10.1016/s0005-2728(97)00046-7. PMID:9332500 doi:http://dx.doi.org/10.1016/s0005-2728(97)00046-7
- ↑ Das A, Silaghi-Dumitrescu R, Ljungdahl LG, Kurtz DM Jr. Cytochrome bd oxidase, oxidative stress, and dioxygen tolerance of the strictly anaerobic bacterium Moorella thermoacetica. J Bacteriol. 2005 Mar;187(6):2020-9. doi: 10.1128/JB.187.6.2020-2029.2005. PMID:15743950 doi:http://dx.doi.org/10.1128/JB.187.6.2020-2029.2005
- ↑ Safarian S, Rajendran C, Muller H, Preu J, Langer JD, Ovchinnikov S, Hirose T, Kusumoto T, Sakamoto J, Michel H. Structure of a bd oxidase indicates similar mechanisms for membrane-integrated oxygen reductases. Science. 2016 Apr 29;352(6285):583-6. doi: 10.1126/science.aaf2477. PMID:27126043 doi:http://dx.doi.org/10.1126/science.aaf2477
- ↑ Safarian S, Rajendran C, Muller H, Preu J, Langer JD, Ovchinnikov S, Hirose T, Kusumoto T, Sakamoto J, Michel H. Structure of a bd oxidase indicates similar mechanisms for membrane-integrated oxygen reductases. Science. 2016 Apr 29;352(6285):583-6. doi: 10.1126/science.aaf2477. PMID:27126043 doi:http://dx.doi.org/10.1126/science.aaf2477
- ↑ Safarian S, Rajendran C, Muller H, Preu J, Langer JD, Ovchinnikov S, Hirose T, Kusumoto T, Sakamoto J, Michel H. Structure of a bd oxidase indicates similar mechanisms for membrane-integrated oxygen reductases. Science. 2016 Apr 29;352(6285):583-6. doi: 10.1126/science.aaf2477. PMID:27126043 doi:http://dx.doi.org/10.1126/science.aaf2477
- ↑ Safarian S, Rajendran C, Muller H, Preu J, Langer JD, Ovchinnikov S, Hirose T, Kusumoto T, Sakamoto J, Michel H. Structure of a bd oxidase indicates similar mechanisms for membrane-integrated oxygen reductases. Science. 2016 Apr 29;352(6285):583-6. doi: 10.1126/science.aaf2477. PMID:27126043 doi:http://dx.doi.org/10.1126/science.aaf2477