This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1616
From Proteopedia
(Difference between revisions)
| Line 13: | Line 13: | ||
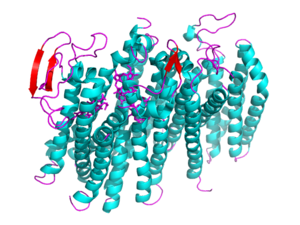
[[Image: 5doq_WHOLE_IMAGE.png|300 px|left|thumb|Figure 2: 6RX4 monomer subunit; alpha helices in teal, beta sheets in purple.]] | [[Image: 5doq_WHOLE_IMAGE.png|300 px|left|thumb|Figure 2: 6RX4 monomer subunit; alpha helices in teal, beta sheets in purple.]] | ||
{{Clear}} | {{Clear}} | ||
| - | This page will be specifically focusing on the structure and overall function of the 6RX4 bd oxidase. 6RX4 is a part of the long(L) quinol-binding domain subfamily that terminal oxidases are classified into. The L-subfamily of bd oxidases are responsible for the survival of acute infectious diseases such as E.Coli and salmonella. The 6RX4's three <scene name='83/832931/Heme/4'>heme</scene> groups, its periplasmically exposed Q loop, and four protein subunits will be of primary focus when identifying the relationship between structure and function. | + | This page will be specifically focusing on the structure and overall function of the 6RX4 bd oxidase. 6RX4 is a part of the long(L) quinol-binding domain subfamily that terminal oxidases are classified into. The L-subfamily of bd oxidases are responsible for the survival of acute infectious diseases such as E.Coli and salmonella. The 6RX4's three <scene name='83/832931/Heme/4'>heme</scene> groups, its periplasmically exposed <scene name='83/832924/Q_loop/3'>Q-loop</scene>, and four protein subunits will be of primary focus when identifying the relationship between structure and function. |
== Function == | == Function == | ||
| Line 23: | Line 23: | ||
== Structural highlights == | == Structural highlights == | ||
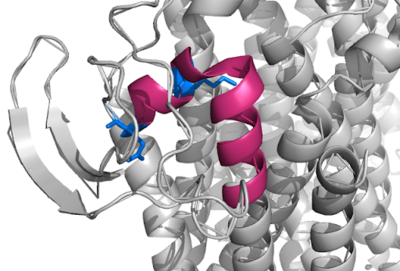
[[Image:Q Loop.png|400 px|right|thumb|Figure 1]] | [[Image:Q Loop.png|400 px|right|thumb|Figure 1]] | ||
| - | This is a sample scene created with SAT to <scene name="/12/3456/Sample/1">color</scene> by Group, and another to make <scene name="/12/3456/Sample/2">a transparent representation</scene> of the protein. You can make your own scenes on SAT starting from scratch or loading and editing one of these sample scenes. | ||
| - | |||
| - | <scene name='83/837229/Residues_22-44_of_bd_oxidase/2'>bd Oxidase Residues 22-44</scene> | ||
Revision as of 04:18, 31 March 2020
bd Oxidase, 5DOQ
| |||||||||||
References
- ↑ 1.0 1.1 Safarian S, Rajendran C, Muller H, Preu J, Langer JD, Ovchinnikov S, Hirose T, Kusumoto T, Sakamoto J, Michel H. Structure of a bd oxidase indicates similar mechanisms for membrane-integrated oxygen reductases. Science. 2016 Apr 29;352(6285):583-6. doi: 10.1126/science.aaf2477. PMID:27126043 doi:http://dx.doi.org/10.1126/science.aaf2477
- ↑ 2.0 2.1 Ransey E, Paredes E, Dey SK, Das SR, Heroux A, Macbeth MR. Crystal structure of the Entamoeba histolytica RNA lariat debranching enzyme EhDbr1 reveals a catalytic Zn(2+) /Mn(2+) heterobinucleation. FEBS Lett. 2017 Jul;591(13):2003-2010. doi: 10.1002/1873-3468.12677. Epub 2017, Jun 14. PMID:28504306 doi:http://dx.doi.org/10.1002/1873-3468.12677
![Figure 1: Overall schematic representation of cytochrome bd; General display of the reduction of molecular oxygen into water [1]](/wiki/images/thumb/e/e6/Proton_graadient.jpg/300px-Proton_graadient.jpg)