This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Harrison L. Smith/Sandbox1
From Proteopedia
(Difference between revisions)
| Line 34: | Line 34: | ||
===Conformation Change=== | ===Conformation Change=== | ||
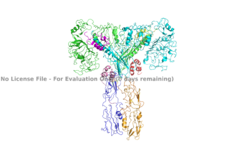
| - | The inactive form of the insulin receptor predominates in low-levels of circulating insulin, whereas the active conformation is seen when insulin binds to any of the 4 receptor sites. The inactive conformation resembles an <scene name='83/832953/Simple_inactivated_receptor/1'>inverted V</scene>, and the active conformation resembles <scene name='83/832953/Ir_dimer_t_state/1'>T shape</scene>. The image of the inverted V conformation shows only a protomer of the inactive alpha subunit because the entire inactive alpha subunit dimer has been unable to be photographed because the transition state has yet to be determined in full. In the V-shape, the FnIII-3 domains are separated by about 120A. At this distance, they cannot work together to autophosphorylate. Upon the binding of insulin to any of the four binding sites, the conformation change will begin and bring the FnIII-3 domains within 40A of each other, which is the T-state conformation. Cryo-EM results have displayed clear representations of FnIII-2 and FnIII-3 domains, but lack in high density results for the transmembrane domain and cannot truly model anything past the two fibronectin domains due to the lack of side chain density. Due to the fact that FnIII-3 is connected to the transmembrane domain and intracellular kinase domains through a short linker, it is suggested that the insulin receptor does extend its T-shape conformation through the cell membrane and into the cell. Therefore, it is expected that the intracellular kinase domains will be in close proximity when this conformation change occurs extracellularly, ultimately allowing for autophosphorylation. <ref> DOI 10.1038/s41467-018-06826-6</ref> | + | The inactive form of the insulin receptor predominates in low-levels of circulating insulin, whereas the active conformation is seen when insulin binds to any of the 4 receptor sites. The inactive conformation resembles an <scene name='83/832953/Simple_inactivated_receptor/1'>inverted V</scene>, and the active conformation resembles <scene name='83/832953/Ir_dimer_t_state/1'>T shape</scene>. The image of the inverted V conformation shows only a protomer of the inactive alpha subunit because the entire inactive alpha subunit dimer has been unable to be photographed because the transition state has yet to be determined in full. In the V-shape, the FnIII-3 domains are separated by about 120A. At this distance, they cannot work together to autophosphorylate. Upon the binding of insulin to any of the four binding sites, the conformation change will begin and bring the FnIII-3 domains within 40A of each other, which is the T-state conformation. Cryo-EM results have displayed clear representations of FnIII-2 and FnIII-3 domains, but lack in high density results for the transmembrane domain and cannot truly model anything past the two fibronectin domains due to the lack of side chain density. Due to the fact that FnIII-3 is connected to the transmembrane domain and intracellular kinase domains through a short linker, it is suggested that the insulin receptor does extend its T-shape conformation through the cell membrane and into the cell. Therefore, it is expected that the intracellular kinase domains will be in close proximity when this conformation change occurs extracellularly, ultimately allowing for autophosphorylation. <ref> DOI 10.1038/s41467-018-06826-6</ref> <ref name="Uchikawa" />''' |
===Binding interactions=== | ===Binding interactions=== | ||
| Line 44: | Line 44: | ||
[https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1010832/ Treatment of Diabetes with Insulin] | [https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1010832/ Treatment of Diabetes with Insulin] | ||
===At the Cellular Level=== | ===At the Cellular Level=== | ||
| - | In the human body, the conformation change from the inactive to active state upon insulin binding has a time constant of six minutes. Once insulin binds and the beta subunits are brought within close proximity, autophosphorylation of the beta subunits begins. Phosphorylation at these sites reaches a maximal level in about one minute, and lasts for approximately six to ten minutes. One insulin receptor substrate has a half-life of 3.5 minutes where it is able to be phosphorylated by the tyrosine kinases of the beta subunit and then act as a central hub to activate further downstream signaling pathways that eventually bring glucose receptors to the surface of the cell to allow for diffusion of glucose into the cell. Once insulin binds to the alpha subunit, the receptor remains active for approximately ten minutes before the insulin is degraded and the receptor returns to its inactive conformation. This time frame puts a perspective on how long it takes for the human body to store excess glucose in their blood stream from a recent meal as glycogen for later use as fuel. | + | In the human body, the conformation change from the inactive to active state upon insulin binding has a time constant of six minutes. Once insulin binds and the beta subunits are brought within close proximity, autophosphorylation of the beta subunits begins. Phosphorylation at these sites reaches a maximal level in about one minute, and lasts for approximately six to ten minutes. One insulin receptor substrate has a half-life of 3.5 minutes where it is able to be phosphorylated by the tyrosine kinases of the beta subunit and then act as a central hub to activate further downstream signaling pathways that eventually bring glucose receptors to the surface of the cell to allow for diffusion of glucose into the cell. Once insulin binds to the alpha subunit, the receptor remains active for approximately ten minutes before the insulin is degraded and the receptor returns to its inactive conformation. This time frame puts a perspective on how long it takes for the human body to store excess glucose in their blood stream from a recent meal as glycogen for later use as fuel. <ref name="Tatulian" />''' |
Revision as of 17:53, 6 April 2020
Homo sapiens Insulin Receptor
| |||||||||||
References
- ↑ 1.0 1.1 1.2 1.3 Tatulian SA. Structural Dynamics of Insulin Receptor and Transmembrane Signaling. Biochemistry. 2015 Sep 15;54(36):5523-32. doi: 10.1021/acs.biochem.5b00805. Epub , 2015 Sep 3. PMID:26322622 doi:http://dx.doi.org/10.1021/acs.biochem.5b00805
- ↑ 2.0 2.1 2.2 Uchikawa E, Choi E, Shang G, Yu H, Bai XC. Activation mechanism of the insulin receptor revealed by cryo-EM structure of the fully liganded receptor-ligand complex. Elife. 2019 Aug 22;8. pii: 48630. doi: 10.7554/eLife.48630. PMID:31436533 doi:http://dx.doi.org/10.7554/eLife.48630
- ↑ Weis F, Menting JG, Margetts MB, Chan SJ, Xu Y, Tennagels N, Wohlfart P, Langer T, Muller CW, Dreyer MK, Lawrence MC. The signalling conformation of the insulin receptor ectodomain. Nat Commun. 2018 Oct 24;9(1):4420. doi: 10.1038/s41467-018-06826-6. PMID:30356040 doi:http://dx.doi.org/10.1038/s41467-018-06826-6
- ↑ Wilcox G. Insulin and insulin resistance. Clin Biochem Rev. 2005 May;26(2):19-39. PMID:16278749
- ↑ Riddle MC. Treatment of diabetes with insulin. From art to science. West J Med. 1983 Jun;138(6):838-46. PMID:6351440
Student Contributors
- Harrison Smith
- Alyssa Ritter