We apologize for Proteopedia being slow to respond. For the past two years, a new implementation of Proteopedia has been being built. Soon, it will replace this 18-year old system. All existing content will be moved to the new system at a date that will be announced here.
Sandbox Reserved 1627
From Proteopedia
(Difference between revisions)
| Line 7: | Line 7: | ||
==Structural Overview== | ==Structural Overview== | ||
| - | The insulin receptor is a [http://en.wikipedia.org/wiki/Dimer_(chemistry) dimer] of heterodimers made of two <scene name='83/832953/Alpha_subunits/4'>alpha subunits</scene> and two <scene name='83/832953/Beta_subunits/3'>beta subunits</scene> <ref name="Tatulian">PMID:26322622</ref>. Within the extracellular ectodomain, there are four potential <scene name='83/832953/Binding_sites/2'>binding sites</scene> that can interact with insulin ligands on the extracellular side of the membrane | + | The insulin receptor is a [http://en.wikipedia.org/wiki/Dimer_(chemistry) dimer] of heterodimers made of two <scene name='83/832953/Alpha_subunits/4'>alpha subunits</scene> and two <scene name='83/832953/Beta_subunits/3'>beta subunits</scene> <ref name="Tatulian">PMID:26322622</ref>. Within the extracellular ectodomain, there are four potential <scene name='83/832953/Binding_sites/2'>binding sites</scene> that can interact with insulin ligands on the extracellular side of the membrane. |
====Alpha Subunits==== | ====Alpha Subunits==== | ||
[[Image:Disulfide bridge between alphas.png|thumb|right|260px|Figure 1: Disulfide bridge (yellow) made of two cysteine residues (blue) that provides a linkage and stability to the two alpha subunits. [http://www.rcsb.org/structure/6sof PDB 6SOF]]] | [[Image:Disulfide bridge between alphas.png|thumb|right|260px|Figure 1: Disulfide bridge (yellow) made of two cysteine residues (blue) that provides a linkage and stability to the two alpha subunits. [http://www.rcsb.org/structure/6sof PDB 6SOF]]] | ||
The alpha subunits make up the extracellular domain ([http://en.wikipedia.org/wiki/Ectodomain ectodomain]) of the insulin receptor and are the sites of insulin binding. The alpha subunit is comprised of two Leucine rich domains (L1 & L2), a Cysteine rich domain (CR), and a C-terminal alpha helix. The alpha and beta subunits are held together by a [http://en.wikipedia.org/wiki/Disulfide disulfide bond] at residue C524 of one alpha subunit, and C524 of the other subunit (Figure1). The actual site of insulin binding occurs at the <scene name='83/832953/Alpha_c_helix/3'>α-CT chain</scene> of one of the sites discussed next and is stabilized by the L1 and L2 domains. Two types of insulin binding sites are present in the alpha subunits, Sites 1 and 1' and then Sites 2 and 2'. Due to structural differences in these binding sites, the first two sites, 1 and 1', have much higher affinity than that of sites 2 and 2'. The sites are in pairs because of the heterodimeric nature of the receptor. Each time an insulin ligand binds to sites 1 and 1', it comes in contact with the L1 domain of one protomer and the alpha-CT chain and FnIII-1 loop of another protomer, which is also known as "cross linking". Insulin can also bind at sites 2 and 2', but the location on the back of the beta sheet of the FnIII-1 domain and lower surface area decreases their binding occupancy. <ref name="Uchikawa"> DOI 10.7554/eLife.48630 </ref>. | The alpha subunits make up the extracellular domain ([http://en.wikipedia.org/wiki/Ectodomain ectodomain]) of the insulin receptor and are the sites of insulin binding. The alpha subunit is comprised of two Leucine rich domains (L1 & L2), a Cysteine rich domain (CR), and a C-terminal alpha helix. The alpha and beta subunits are held together by a [http://en.wikipedia.org/wiki/Disulfide disulfide bond] at residue C524 of one alpha subunit, and C524 of the other subunit (Figure1). The actual site of insulin binding occurs at the <scene name='83/832953/Alpha_c_helix/3'>α-CT chain</scene> of one of the sites discussed next and is stabilized by the L1 and L2 domains. Two types of insulin binding sites are present in the alpha subunits, Sites 1 and 1' and then Sites 2 and 2'. Due to structural differences in these binding sites, the first two sites, 1 and 1', have much higher affinity than that of sites 2 and 2'. The sites are in pairs because of the heterodimeric nature of the receptor. Each time an insulin ligand binds to sites 1 and 1', it comes in contact with the L1 domain of one protomer and the alpha-CT chain and FnIII-1 loop of another protomer, which is also known as "cross linking". Insulin can also bind at sites 2 and 2', but the location on the back of the beta sheet of the FnIII-1 domain and lower surface area decreases their binding occupancy. <ref name="Uchikawa"> DOI 10.7554/eLife.48630 </ref>. | ||
===Beta Subunits=== | ===Beta Subunits=== | ||
| - | The beta subunit spans from the extracellular domain across the transmembrane region and into the intracellular portion of the insulin receptor. The beta subunit is composed of part of [http://en.wikipedia.org/wiki/Fibronectin fibronectin] domain III-2 and all of Fibronectin domain III-3. The beta subunit's FnIII-3 domain has links through the transmembrane region into the intracellular part of the membrane | + | The beta subunit spans from the extracellular domain across the transmembrane region and into the intracellular portion of the insulin receptor. The beta subunit is composed of part of [http://en.wikipedia.org/wiki/Fibronectin fibronectin] domain III-2 and all of Fibronectin domain III-3. The beta subunit's FnIII-3 domain has links through the transmembrane region into the intracellular part of the membrane. |
===Cryo-EM Structural Imaging=== | ===Cryo-EM Structural Imaging=== | ||
| Line 25: | Line 25: | ||
===Conformation Change=== | ===Conformation Change=== | ||
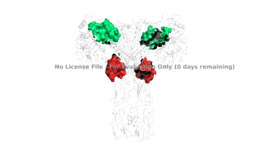
| - | Structures of the inactive inverted V conformation only contains a single <scene name='83/832953/Inactive_insulin_receptor/3'>protomer of the extracellular alpha and beta subunits</scene> because the entire inactive alpha subunit dimer has been unable to be photographed because the transition state has yet to be determined in full. In the V-shape, the FnIII-3 domains are separated by about 120Å which keeps the tyrosine kinase domains separated. In the V-shape, autophosphorylation and downstream signaling cannot be initiated. Upon the binding of insulin to either the 1 or 1' site, the conformation change will begin and bring the FnIII-3 domains within 40Å of each other | + | Structures of the inactive inverted V conformation only contains a single <scene name='83/832953/Inactive_insulin_receptor/3'>protomer of the extracellular alpha and beta subunits</scene> because the entire inactive alpha subunit dimer has been unable to be photographed because the transition state has yet to be determined in full. In the V-shape, the FnIII-3 domains are separated by about 120Å which keeps the tyrosine kinase domains separated. In the V-shape, autophosphorylation and downstream signaling cannot be initiated. Upon the binding of insulin to either the 1 or 1' site, the conformation change will begin and bring the FnIII-3 domains within 40Å of each other to induce the T-state conformation. <ref> DOI 10.1038/s41467-018-06826-6</ref> <ref name="Uchikawa" /> The T shape conformation is well observed in the alpha subunit. It is horizontally composed of L1, CR (including the alpha-CT chain), and L2 domains and vertically composed of the FnIII-1, 2, and 3 domains. This structural transition will facilitate the autophosphorylation of the tyrosine kinase domain. |
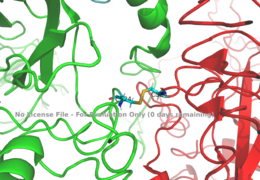
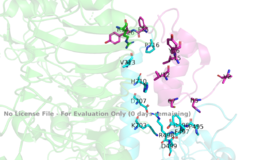
===Binding interactions=== | ===Binding interactions=== | ||
Revision as of 21:45, 16 April 2020
Homo sapiens Insulin Receptor
| |||||||||||
References
- ↑ 1.0 1.1 De Meyts P. The Insulin Receptor and Its Signal Transduction Network PMID:27512793
- ↑ 2.0 2.1 2.2 Tatulian SA. Structural Dynamics of Insulin Receptor and Transmembrane Signaling. Biochemistry. 2015 Sep 15;54(36):5523-32. doi: 10.1021/acs.biochem.5b00805. Epub , 2015 Sep 3. PMID:26322622 doi:http://dx.doi.org/10.1021/acs.biochem.5b00805
- ↑ 3.0 3.1 3.2 Uchikawa E, Choi E, Shang G, Yu H, Bai XC. Activation mechanism of the insulin receptor revealed by cryo-EM structure of the fully liganded receptor-ligand complex. Elife. 2019 Aug 22;8. pii: 48630. doi: 10.7554/eLife.48630. PMID:31436533 doi:http://dx.doi.org/10.7554/eLife.48630
- ↑ Weis F, Menting JG, Margetts MB, Chan SJ, Xu Y, Tennagels N, Wohlfart P, Langer T, Muller CW, Dreyer MK, Lawrence MC. The signalling conformation of the insulin receptor ectodomain. Nat Commun. 2018 Oct 24;9(1):4420. doi: 10.1038/s41467-018-06826-6. PMID:30356040 doi:http://dx.doi.org/10.1038/s41467-018-06826-6
- ↑ Uchikawa E, Choi E, Shang G, Yu H, Bai XC. Activation mechanism of the insulin receptor revealed by cryo-EM structure of the fully liganded receptor-ligand complex. Elife. 2019 Aug 22;8. pii: 48630. doi: 10.7554/eLife.48630. PMID:31436533 doi:http://dx.doi.org/10.7554/eLife.48630
- ↑ Boucher J, Kleinridders A, Kahn CR. Insulin receptor signaling in normal and insulin-resistant states. Cold Spring Harb Perspect Biol. 2014 Jan 1;6(1). pii: 6/1/a009191. doi:, 10.1101/cshperspect.a009191. PMID:24384568 doi:http://dx.doi.org/10.1101/cshperspect.a009191
- ↑ Wilcox G. Insulin and insulin resistance. Clin Biochem Rev. 2005 May;26(2):19-39. PMID:16278749
- ↑ Riddle MC. Treatment of diabetes with insulin. From art to science. West J Med. 1983 Jun;138(6):838-46. PMID:6351440
Student Contributors
- Harrison Smith
- Alyssa Ritter