This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Felipe de Melo Santana/Sandbox 1
From Proteopedia
| Line 14: | Line 14: | ||
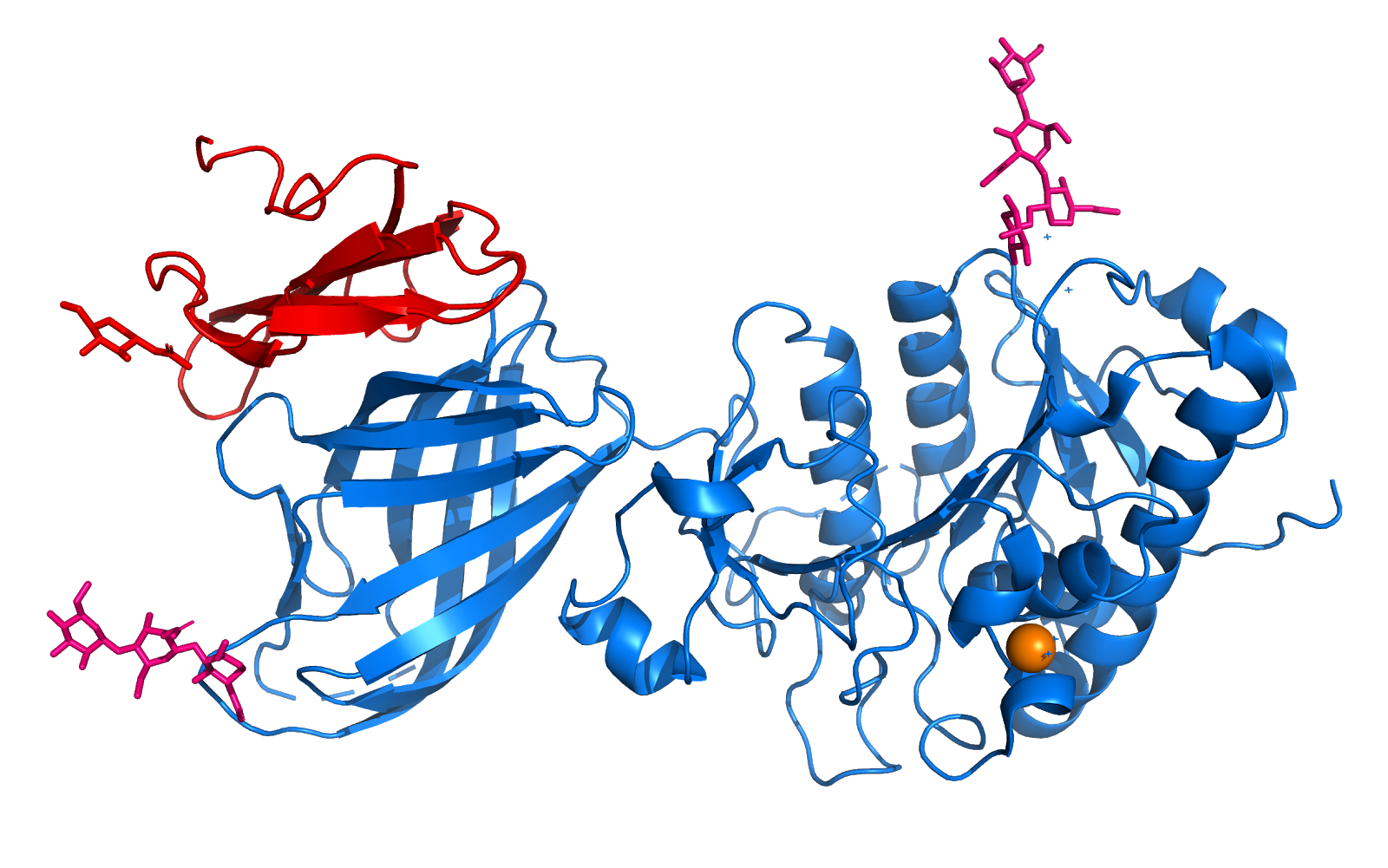
The remaining C-terminal 30 Structure Determination residues come back to domain I to form an edge of the domain . The two domains exhibit a similar folding pattern consisting of a central b sheet flanked by helices and loops. | The remaining C-terminal 30 Structure Determination residues come back to domain I to form an edge of the domain . The two domains exhibit a similar folding pattern consisting of a central b sheet flanked by helices and loops. | ||
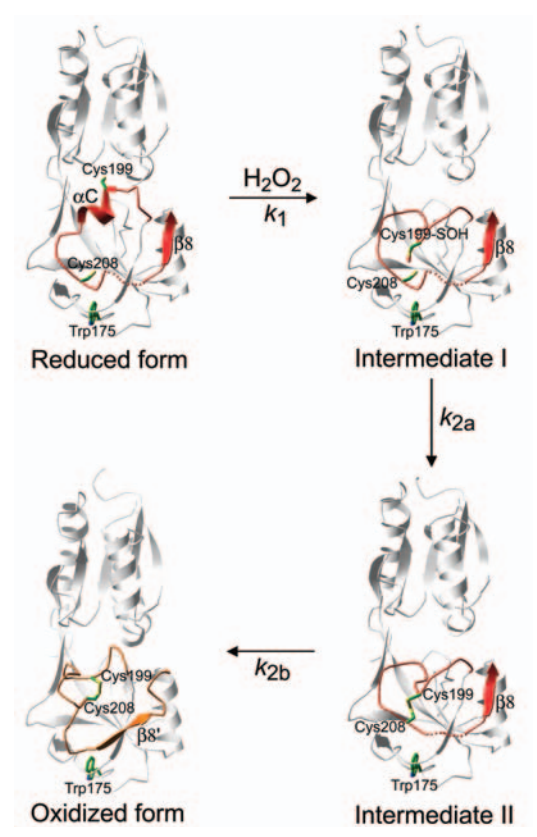
The oxidized form is characterized by the ediction of the domain II, resulting in a significant rearrangement of secondary structures. | The oxidized form is characterized by the ediction of the domain II, resulting in a significant rearrangement of secondary structures. | ||
| + | |||
| + | [[Image:PHOTO-2020-06-20-17-03-18.jpg]] | ||
Revision as of 20:09, 20 June 2020
1I69
| |||||||||||
References
Zheng, M., Aslund, F., and Storz, G. (1998). Activation of the OxyR transcription factor by reversible disulfide bond formation. Science 279, 1718–1721.
Choi H, Kim S, Mukhopadhyay P, et al. Structural basis of the redox switch in the OxyR transcription factor. Cell. 2001;105(1):103-113. doi:10.1016/s0092-8674(01)00300-2
Toledano, M.B., Kullik, I., Trinh, F., Baird, P.T., Schneider, T.D., and Storz, G. (1994). Redox-dependent shift of OxyR-DNA contacts along an extended DNA-binding site: a mechanism for differential promoter selection. Cell 78, 897–909.
Storz, G., Tartaglia, L.A., and Ames, B.N. (1990). Transcriptional regulator of oxidative stress-inducible genes: direct activation by oxidation. Science 248, 189–194
Kullik, I., Toledano, M.B., Tartaglia, L.A., and Storz, G. (1995a). Mutational analysis of the redox-sensitive transcriptional regulator OxyR: regions important for oxidation and transcriptional activation. J. Bacteriol. 177, 1275–1284.
Lee C, Lee SM, Mukhopadhyay P, et al. Redox regulation of OxyR requires specific disulfide bond formation involving a rapid kinetic reaction path. Nat Struct Mol Biol. 2004;11(12):1179-1185. doi:10.1038/nsmb856
Previato, M. Regulation of genes involved in oxidative stress response in Caulobacter crescentus. 2013. 134 p. Masters thesis (Microbiology) – Instituto de Ciências Biomédicas, Universidade de São Paulo, São Paulo, 2013.
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644