We apologize for Proteopedia being slow to respond. For the past two years, a new implementation of Proteopedia has been being built. Soon, it will replace this 18-year old system. All existing content will be moved to the new system at a date that will be announced here.
User:Andre Wu Le Chun/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 9: | Line 9: | ||
6svb is a homotrimer transmembrane glycoprotein(S) protruding from the viral surface¹. It is composed by two subunits that are fundamental for the virus entry in the cell: The S1 subunit, which contains the receptor binding domain (RBD), is responsible for binding to the host cell receptor, and the S2 subunit, responsible for fusion of the viral and cellular membranes. In order to allow the binding of the RBD in S1 with the host cell, the spike protein goes through conformational changes that start from a pre-fusion conformation to a stable post-fusion conformation, in which the receptor-binding domain assumes a up conformation. | 6svb is a homotrimer transmembrane glycoprotein(S) protruding from the viral surface¹. It is composed by two subunits that are fundamental for the virus entry in the cell: The S1 subunit, which contains the receptor binding domain (RBD), is responsible for binding to the host cell receptor, and the S2 subunit, responsible for fusion of the viral and cellular membranes. In order to allow the binding of the RBD in S1 with the host cell, the spike protein goes through conformational changes that start from a pre-fusion conformation to a stable post-fusion conformation, in which the receptor-binding domain assumes a up conformation. | ||
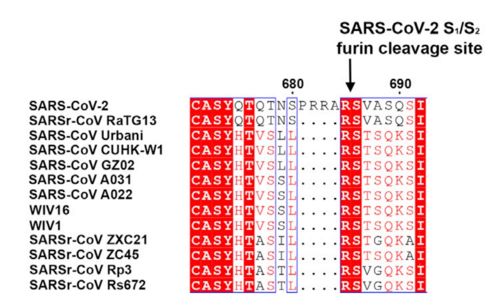
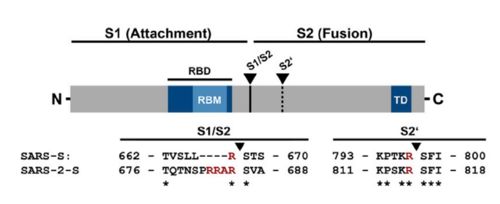
| - | This protein also shares amino acid sequence identity of 76% with SARS-CoV, however, as shown | + | <scene name='84/844931/S_receptor-binding_domain/1'>Text To Be Displayed</scene> |
| + | |||
| + | This protein also shares amino acid sequence identity of 76% with SARS-CoV, however, as shown in the figure bellow, it has inserted amino acids that indicate a furin like cleavage site in the S1/S2 boundary. that must be primed in order to enable the viral pathogenicity. In the Receptor binding domain, units of beta-sheets, alpha-helices and loops can be observed. | ||
| Line 37: | Line 39: | ||
¹https://www.sciencedirect.com/science/article/pii/S0065352719300284 | ¹https://www.sciencedirect.com/science/article/pii/S0065352719300284 | ||
²https://www.biorxiv.org/content/10.1101/2020.02.19.956581v1.full | ²https://www.biorxiv.org/content/10.1101/2020.02.19.956581v1.full | ||
| + | https://www.sciencedirect.com/science/article/pii/S0092867420302294 | ||
<references/> | <references/> | ||
Revision as of 19:31, 21 June 2020
6vsb
Prefusion 2019-nCoV spike glycoprotein with a single receptor-binding domain up
| |||||||||||
References
¹https://www.sciencedirect.com/science/article/pii/S0065352719300284 ²https://www.biorxiv.org/content/10.1101/2020.02.19.956581v1.full https://www.sciencedirect.com/science/article/pii/S0092867420302294
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644