This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Green Fluorescent Protein
From Proteopedia
| Line 46: | Line 46: | ||
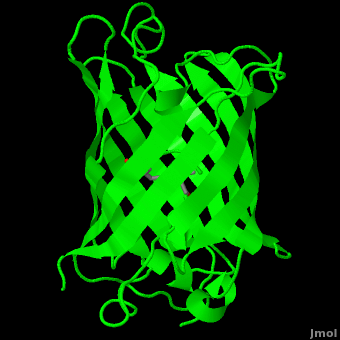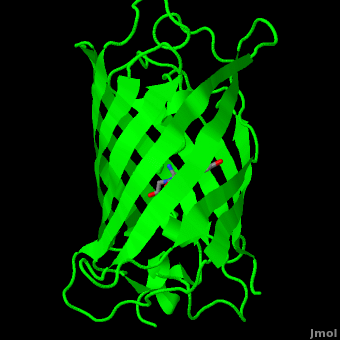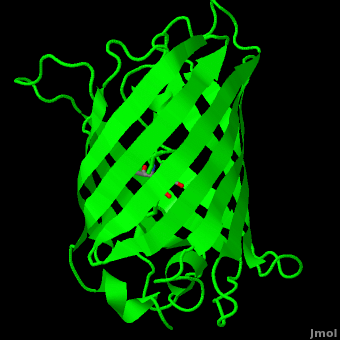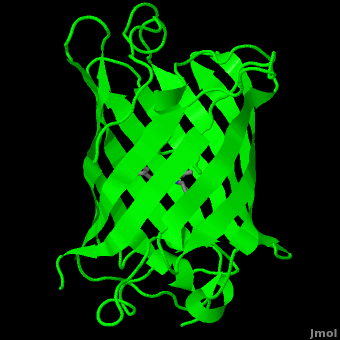
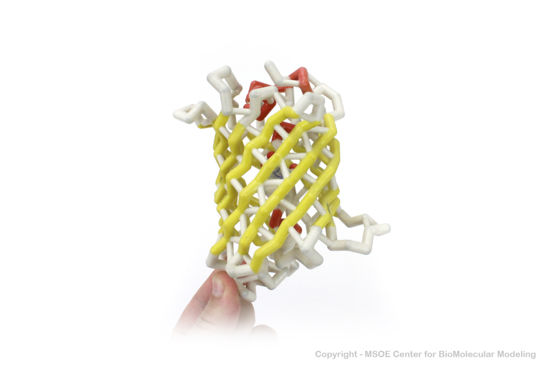
The <scene name='10/100139/Chromophore/2'>chromophore</scene> (<scene name='10/100139/Green_fluorescent_protein/1'>top view</scene>) of GFP is located at the center of the β-barrel with a wild-type excitation peak of 395 nm, and a minor peak at 475 nm (about three times less intense<ref name="Tsien" />) <ref name="Yang" /><ref name="Cubitt" /><ref name="Ormo" /><ref name="Phillips" /> with extinction coefficients of approximately 30,000 and 7,000 M<sup>-1</sup> cm<sup>-1</sup>, respectively.<ref name="Yang" /><ref name="Phillips" /> Interestingly, the ''Aequorea victoria'' jellyfish utilizes the smaller of the two excitation peaks as pure aequorin emits a light of 470 nm.<ref name="Tsien">Tsien, Roger Y. 1998. The Green Fluorescent Protein. Annual Review in Biochemistry. 67:509-544.</ref> The relative amplitudes of these two excitation peaks can vary depending on environmental factors and previous illumination.<ref name="Ormo" /> For example, continued excitation leads to a diminution of the 395 nm excitation peak with a reciprocal amplification of the 475 nm peak.<ref name="Phillips" /> Regardless of absorption, the chromophore of GFP emits light of 508 nm.<ref name="Yang" /><ref name="Cubitt" /><ref name="Ormo" /><ref name="Phillips" /> | The <scene name='10/100139/Chromophore/2'>chromophore</scene> (<scene name='10/100139/Green_fluorescent_protein/1'>top view</scene>) of GFP is located at the center of the β-barrel with a wild-type excitation peak of 395 nm, and a minor peak at 475 nm (about three times less intense<ref name="Tsien" />) <ref name="Yang" /><ref name="Cubitt" /><ref name="Ormo" /><ref name="Phillips" /> with extinction coefficients of approximately 30,000 and 7,000 M<sup>-1</sup> cm<sup>-1</sup>, respectively.<ref name="Yang" /><ref name="Phillips" /> Interestingly, the ''Aequorea victoria'' jellyfish utilizes the smaller of the two excitation peaks as pure aequorin emits a light of 470 nm.<ref name="Tsien">Tsien, Roger Y. 1998. The Green Fluorescent Protein. Annual Review in Biochemistry. 67:509-544.</ref> The relative amplitudes of these two excitation peaks can vary depending on environmental factors and previous illumination.<ref name="Ormo" /> For example, continued excitation leads to a diminution of the 395 nm excitation peak with a reciprocal amplification of the 475 nm peak.<ref name="Phillips" /> Regardless of absorption, the chromophore of GFP emits light of 508 nm.<ref name="Yang" /><ref name="Cubitt" /><ref name="Ormo" /><ref name="Phillips" /> | ||
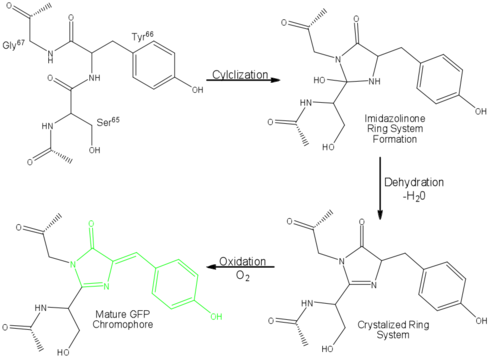
| - | Three amino residues in the central α-helix constitute the fluorophore of GFP: Ser<sup>65</sup>Tyr<sup>66</sup>Gly<sup>67</sup> (see below) or of EGFP: of GFP: Thr<sup>65</sup>Tyr<sup>66</sup>Gly<sup>67</sup>. Tsien et al. discovered that this tri-peptide sequence is post-translationally modified by internal cyclization and oxidation<ref name="Haldar" /> to produce a <scene name='10/100139/Chromophore_structure/3'>4-(p-hydroxybenzylidene)-imidazolidin-5-one</scene> structure (highlight atoms from <jmol><jmolLink><script> select (*.C1, *.CA1, *.N1, *.CB1, *.CG1, *.OG1) and 66; selectionHalos ON; delay 1.5;selectionHalos OFF;</script><text>⚞Thr 65⚟</text></jmolLink> </jmol>, <jmol><jmolLink><script> select (*.C2, *.O2, *.CA2, *.N2, *.CB2, *.CG2, *.CD1, *.CD2, *.CE1, *.CE2, *.CZ, *.OH) and 66; selectionHalos ON; delay 1.5;selectionHalos OFF;</script><text>⚞Tyr 66⚟</text></jmolLink> </jmol>, <jmol><jmolLink><script> select (*.O3,*.C3,*.CA3,*.N3) and 66; selectionHalos ON; delay 1.5;selectionHalos OFF;</script><text>⚞Gly 67⚟</text></jmolLink></jmol>).<ref name="Yang" /> Studies with E. coli proposed a sequential mechanism for the formation of the fluorophore that was initiated by a rapid cyclization between Ser<sup>65</sup> and Gly<sup>67</sup> to form an imidazolin-5-one intermediate.<ref name="Yang" /> This rapid cyclization is carried out via nucleophilic attack of the amino group from Gly<sup>67</sup> on the carbonyl group of Ser<sup>65</sup> to form a five-membered ring. The loss of water then forms the imidazolin-5-one intermediate.<ref name="Cubitt" /> Cyclization is succeeded by a much slower rate-limiting oxygenation of the Tyr<sup>66</sup> hydroxybenzyl side chain by atmospheric oxygen (No fluorescence was seen in anaerobically grown E. coli.), resulting in the 4-(p-hydroxybenzylidene)-imidazolidin-5-one stucture.<ref name="Yang" /><ref name="Cubitt" /><ref name="Phillips" /> The double bond that results from this series of reactions results in the linkage of the two π-systems of the rings, forming a <scene name='10/100139/Chromophore/ | + | Three amino residues in the central α-helix constitute the fluorophore of GFP: Ser<sup>65</sup>Tyr<sup>66</sup>Gly<sup>67</sup> (see below) or of EGFP: of GFP: Thr<sup>65</sup>Tyr<sup>66</sup>Gly<sup>67</sup>. Tsien et al. discovered that this tri-peptide sequence is post-translationally modified by internal cyclization and oxidation<ref name="Haldar" /> to produce a <scene name='10/100139/Chromophore_structure/3'>4-(p-hydroxybenzylidene)-imidazolidin-5-one</scene> structure (highlight atoms from <jmol><jmolLink><script> select (*.C1, *.CA1, *.N1, *.CB1, *.CG1, *.OG1) and 66; selectionHalos ON; delay 1.5;selectionHalos OFF;</script><text>⚞Thr 65⚟</text></jmolLink> </jmol>, <jmol><jmolLink><script> select (*.C2, *.O2, *.CA2, *.N2, *.CB2, *.CG2, *.CD1, *.CD2, *.CE1, *.CE2, *.CZ, *.OH) and 66; selectionHalos ON; delay 1.5;selectionHalos OFF;</script><text>⚞Tyr 66⚟</text></jmolLink> </jmol>, <jmol><jmolLink><script> select (*.O3,*.C3,*.CA3,*.N3) and 66; selectionHalos ON; delay 1.5;selectionHalos OFF;</script><text>⚞Gly 67⚟</text></jmolLink></jmol>).<ref name="Yang" /> Studies with E. coli proposed a sequential mechanism for the formation of the fluorophore that was initiated by a rapid cyclization between Ser<sup>65</sup> and Gly<sup>67</sup> to form an imidazolin-5-one intermediate.<ref name="Yang" /> This rapid cyclization is carried out via nucleophilic attack of the amino group from Gly<sup>67</sup> on the carbonyl group of Ser<sup>65</sup> to form a five-membered ring. The loss of water then forms the imidazolin-5-one intermediate.<ref name="Cubitt" /> Cyclization is succeeded by a much slower rate-limiting oxygenation of the Tyr<sup>66</sup> hydroxybenzyl side chain by atmospheric oxygen (No fluorescence was seen in anaerobically grown E. coli.), resulting in the 4-(p-hydroxybenzylidene)-imidazolidin-5-one stucture.<ref name="Yang" /><ref name="Cubitt" /><ref name="Phillips" /> The double bond that results from this series of reactions results in the linkage of the two π-systems of the rings, forming a <scene name='10/100139/Chromophore/5'>larger conjugated system</scene> essential for fluorophore stability. <ref name="Bublitz"> Bublitz G, King BA, Boxer SG. 1998. Electronic structure of the chromophore in green fluorescent protein (GFP). Journal of the American Chemical Society. 120(36): 9370-9371. DOI 10.1021/ja98160e.</ref> |
[[Image:GFP Chromophore.png|center|489x360px]] | [[Image:GFP Chromophore.png|center|489x360px]] | ||
Revision as of 13:04, 27 October 2020
| |||||||||||
Reference for this Structure
Ormo M, Cubitt AB, Kallio K, Gross LA, Tsien RY, Remington SJ. 1996. Crystal structure of the Aequorea victoria green fluorescent protein. Science. 273(5280):1392-1395. DOI 10.1126/science.273.5280.1392.
References
- ↑ 1.0 1.1 [1], Protein Database (PDBsum): 1ema. European Bioinformatics (EBI); 2009.
- ↑ 2.00 2.01 2.02 2.03 2.04 2.05 2.06 2.07 2.08 2.09 2.10 2.11 2.12 2.13 2.14 2.15 2.16 2.17 [2], Yang F, Moss LG, Phillips GN Jr. 1996. The molecular structure of green fluorescent protein. Biotechnology. 14: 1246-1251. DOI 10.1038/nbt1096-1246.
- ↑ 3.0 3.1 3.2 3.3 3.4 3.5 Tsien, Roger Y. 1998. The Green Fluorescent Protein. Annual Review in Biochemistry. 67:509-544.
- ↑ 4.0 4.1 4.2 4.3 4.4 4.5 4.6 4.7 [3], Haldar S, Chattopadhyay A. 2009. The green journey. J Fluoresc. 19:1-2. DOI 10.1007/s10895-008-0455-6; biographical background on Douglas Prasher, Martin Chalfie and Roger Tsien.
- ↑ 5.0 5.1 5.2 5.3 [4], Shimomura O. The discovery of green fluorescent protein. Nobel Prize Lecture; 2009;; biographical background at Wikipedia.
- ↑ [5],Cowles D, Cowles J. Aequorea victoria. 2007. Walla Wall University.
- ↑ Primary structure at www.ebi.aci.uk.
- ↑ 8.00 8.01 8.02 8.03 8.04 8.05 8.06 8.07 8.08 8.09 8.10 8.11 8.12 8.13 Ormo M, Cubitt AB, Kallio K, Gross LA, Tsien RY, Remington SJ. 1996. Crystal structure of the Aequorea victoria green fluorescent protein. Science. 273(5280):1392-1395. DOI 10.1126/science.273.5280.1392.
- ↑ 9.00 9.01 9.02 9.03 9.04 9.05 9.06 9.07 9.08 9.09 9.10 9.11 9.12 9.13 Phillips GN Jr. Structure and dynamics of green fluorescent protein. Curr Opin Struct Biol. 1997 Dec;7(6):821-7. PMID:9434902
- ↑ Andrews BT, Gosavi S, Finke JM, Onuchic JN, Jennings PA. The dual-basin landscape in GFP folding. Proc Natl Acad Sci U S A. 2008 Aug 26;105(34):12283-8. Epub 2008 Aug 19. PMID:18713871
- ↑ 11.0 11.1 11.2 11.3 11.4 11.5 [6],Cubitt AB, Heim R, Adams SR, Boyd AE, Gross LA, Tsien R. 1995. Understanding, improving, and using green fluorescent protein. Trends in Biochemical Sciences. 20(11): 448-455. DOI 0.1016/S0968-0004(00)89099-4.
- ↑ Bublitz G, King BA, Boxer SG. 1998. Electronic structure of the chromophore in green fluorescent protein (GFP). Journal of the American Chemical Society. 120(36): 9370-9371. DOI 10.1021/ja98160e.
- ↑ van Thor JJ, Sage, JT. 2006. Charge transfer in green fluorescent protein. Photochemical & Photobiological Sciences. 5:597-602. DOI 10.1039/b516525c.
- ↑ 14.0 14.1 Lammich L, Petersen MA, Nielsen MB, Andersen LH. The gas-phase absorption spectrum of a neutral GFP model chromophore. Biophys J. 2007 Jan 1;92(1):201-7. Epub 2006 Oct 13. PMID:17040991 doi:10.1529/biophysj.106.093674
- ↑ Information about edge-face (CH/π) interactions.
- ↑ Fang C, Frontiera RR, Tran R, Mathies RA. Mapping GFP structure evolution during proton transfer with femtosecond Raman spectroscopy. Nature. 2009 Nov 12;462(7270):200-4. PMID:19907490 doi:10.1038/nature08527
Additional Resources
- For additional information, see: Colored & Bioluminescent Proteins
- First Glance
- PDBsum: 1ema
- RCSB PDB 1ema
- OCA
- UniProt: P42212
- Scop: P42212
- CATH: 1emaA00
- Pfam: PF01353
- InterPro: IPR000786
- GFP featured at the Molecule of the Month series of tutorials by David Goodsell.
- Inside green fluorescent protein - editor's summary that accompanied structural detail of GFP chromophore on the cover of Nature.
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Wayne Decatur, Karsten Theis, Eran Hodis, Laura Carbone, Karl Oberholser, Mark Hoelzer, Joel L. Sussman, Alexander Berchansky, Jaime Prilusky, Joseph M. Steinberger, David Canner