This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox GGC1
From Proteopedia
| Line 9: | Line 9: | ||
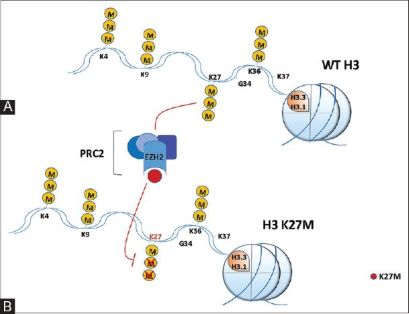
There have been studies that have identified mutations encoding a K27M substitution and there have also been mutations that encoded | There have been studies that have identified mutations encoding a K27M substitution and there have also been mutations that encoded | ||
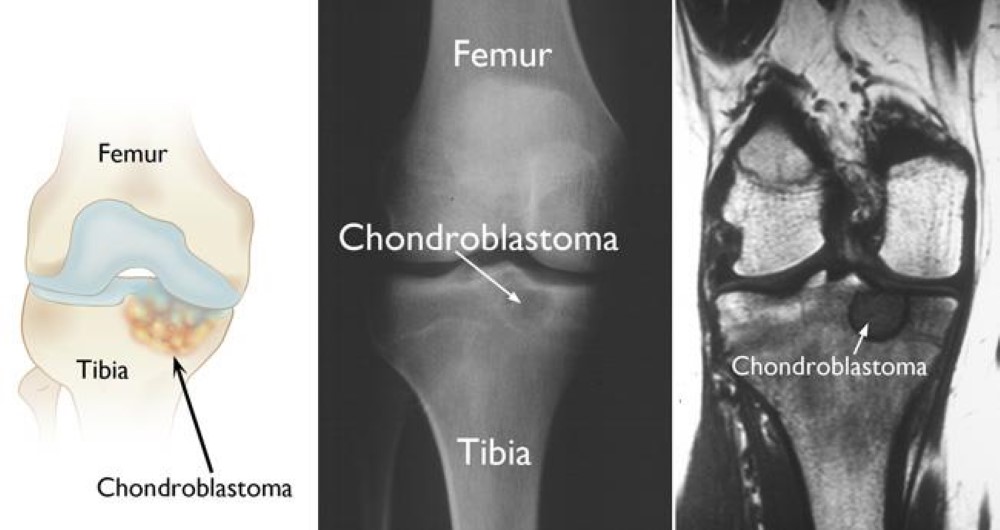
GLY 34 to ARG or VAL called the G34R/V substitution. K27M tumors are present in ex: spinal cord, thalamus, pons, brainstem and G34R/V tumors are shown in the cerebral hemispheres. There are mutations in H3.3 that are found in different types of bone tumors like [https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4446520/ chrondroblastoma] for example and giant cell tumors of the bone. [https://cancerdiscovery.aacrjournals.org/content/3/12/1329.1 Chondroblastoma]arises in children and in young adults in the cartilage of the growth plates of the long bones and is most typically benign. | GLY 34 to ARG or VAL called the G34R/V substitution. K27M tumors are present in ex: spinal cord, thalamus, pons, brainstem and G34R/V tumors are shown in the cerebral hemispheres. There are mutations in H3.3 that are found in different types of bone tumors like [https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4446520/ chrondroblastoma] for example and giant cell tumors of the bone. [https://cancerdiscovery.aacrjournals.org/content/3/12/1329.1 Chondroblastoma]arises in children and in young adults in the cartilage of the growth plates of the long bones and is most typically benign. | ||
| + | |||
[[Image:Glioma_2018_1_4_117_240231_f1.jpg]] | [[Image:Glioma_2018_1_4_117_240231_f1.jpg]] | ||
Revision as of 13:39, 16 November 2020
Histone H3.3
| |||||||||||
References
1. Arimura, Y.; Shirayama, K.; Horikoshi, N.; Fujita, R.; Taguchi, H.; Kagawa, W.; Fukagawa, T.; Almouzni, G.; Kurumizaka, H. Crystal structure and stable property of the cancer-associated heterotypic nucleosome containing CENP-A and H3.3. https://www.nature.com/articles/srep07115 (accessed Nov 1, 2020).
2. Cancer Discovery Science Writers. Histone H3.3 Mutations Are Cancer Type-Specific. https://cancerdiscovery.aacrjournals.org/content/3/12/1329.1 (accessed Nov 14, 2020).
3. Gianno, F.; Antonelli, M.; Ferretti2018, E.; Massimino, M.; Arcella, A.; Giangaspero, F. Pediatric high-grade glioma: A heterogeneous group of neoplasms with different molecular drivers.  (accessed Nov 16, 2020).
(accessed Nov 16, 2020).
4. Kallappagoudar, S.; Yadav, R. K.; Lowe, B. R.; Partridge, J. F. Histone H3 mutations--a special role for H3.3 in tumorigenesis? https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4446520/ (accessed Nov 1, 2020).
5. Morell, N.; Rajani, R. Chondroblastoma - OrthoInfo - AAOS. https://orthoinfo.aaos.org/en/diseases--conditions/chondroblastoma (accessed Nov 16, 2020).
6.panelRuiGuo111LijuanZheng111Juw WonPark2RuituLv1HaoChen1FangfangJiao1WenqiXu1ShirongMu3HongWen45JinsongQiu6ZhentianWang1PengyuanYang1FeizhenWu1JingyiHui3XiangdongFu6XiaobingShi4512Yujiang GenoShi7812YiXing212…YangShi891012, A. links open overlay; RuiGuo111; 1; 11; LijuanZheng111; Juw WonPark2; 2; RuituLv1; HaoChen1; FangfangJiao1; WenqiXu1; ShirongMu3; 3; HongWen45; 4; 5; JinsongQiu6; 6; ZhentianWang1; PengyuanYang1; FeizhenWu1; JingyiHui3; XiangdongFu6; XiaobingShi4512; 12; Yujiang GenoShi7812; 7; 8; YiXing212; YangShi891012; 9; 10; Highlights•BS69/ZMYND11 binds H3.3K36me3 and colocalizes with H3.3K36me3 in gene bodies•BS69 directly interacts with EFTUD2; SummaryBS69 (also called ZMYND11) contains tandemly arranged PHD. BS69/ZMYND11 Reads and Connects Histone H3.3 Lysine 36 Trimethylation-Decorated Chromatin to Regulated Pre-mRNA Processing. https://reader.elsevier.com/reader/sd/pii/S1097276514006777?token=A4FD3B8CDE2F310EA514C66E96DC4489F79C8EA96F6FC878DCD4BFC066FA809C2E83C8A9B57353A53915171AD2491D4C (accessed Nov 16, 2020).
7. UniProt ConsortiumEuropean Bioinformatics InstituteProtein Information ResourceSIB Swiss Institute of Bioinformatics. Histone H3.3. https://www.uniprot.org/uniprot/P84243 (accessed Nov 1, 2020).
8.Yuen, B. T. K.; Knoepfler, P. S. Histone H3.3 mutations: a variant path to cancer. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3882088/ (accessed Nov 16, 2020).