This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Tachyplesin
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
<StructureSection load='' size='340' side='right' caption='Tacyplesin I (PDB code [[1ma2]])' scene='67/671725/First_scene/2'> | <StructureSection load='' size='340' side='right' caption='Tacyplesin I (PDB code [[1ma2]])' scene='67/671725/First_scene/2'> | ||
== Introduction == | == Introduction == | ||
| - | Tachyplesin I | + | '''Tachyplesin I, II and III''' are [http://en.wikipedia.org/wiki/Antimicrobial_peptides antimicrobial polypeptide] originally detected in the leukocytes of Japanese [http://en.wikipedia.org/wiki/Horseshoe_crab Horse Shoe Crab]. It has been reported to inhibit the growth of [http://en.wikipedia.org/wiki/Bacteria bacteria], [http://en.wikipedia.org/wiki/Fungus fungui] and [http://en.wikipedia.org/wiki/Virus viruses]. |
The antimicrobial activity of the polypeptide is contributed by electrostatic interaction between the negatively charged membrane of bacteria and fungi to positively charged part of <scene name='67/671725/Cationic_peptide_tpi/3'> TP-I </scene> <ref name=Laederach>PMID:12369825</ref> (see the {{Template:ColorKey_Hydrophobic}} and {{Template:ColorKey_Charge_Cationic}} amino acids). | The antimicrobial activity of the polypeptide is contributed by electrostatic interaction between the negatively charged membrane of bacteria and fungi to positively charged part of <scene name='67/671725/Cationic_peptide_tpi/3'> TP-I </scene> <ref name=Laederach>PMID:12369825</ref> (see the {{Template:ColorKey_Hydrophobic}} and {{Template:ColorKey_Charge_Cationic}} amino acids). | ||
Specifically, TP-I shows high affinity for negatively charged [http://en.wikipedia.org/wiki/Lipopolysaccharide lipopolysaccharides (LPS)] of [http://en.wikipedia.org/wiki/Gram-negative_bacteria gram-negative bacteria], thus neutralizing its effects. | Specifically, TP-I shows high affinity for negatively charged [http://en.wikipedia.org/wiki/Lipopolysaccharide lipopolysaccharides (LPS)] of [http://en.wikipedia.org/wiki/Gram-negative_bacteria gram-negative bacteria], thus neutralizing its effects. | ||
| Line 14: | Line 14: | ||
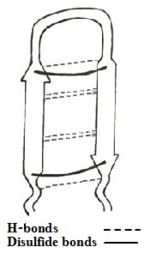
[http://en.wikipedia.org/wiki/Nuclear_magnetic_resonance NMR] studies have shown that TP-I undergoes a conformational change from <scene name='67/671725/First_scene/5'>water surrounding</scene> to <scene name='67/671725/Tp_i_in_the_presence_of_lps/4'>presence of LPS</scene>, making it <scene name='67/671725/Conformation_change/16'>more rigid and twisted</scene> than in the presence of water<ref name=Kushibiki>PMID:24389234</ref>. Moreover a docking model suggests the stability of the structure of TP-I is increased in the presence of LPS by the binding of the N and C termini of TP-I to LPS. The conformational change of TP-I seems to be crucial for its antimicrobial activity, since rearrangement of TP-I structure makes it more amphiphilic to negatively charged membrane of bacteria and fungus<ref name=Laederach>PMID:12369825</ref>. | [http://en.wikipedia.org/wiki/Nuclear_magnetic_resonance NMR] studies have shown that TP-I undergoes a conformational change from <scene name='67/671725/First_scene/5'>water surrounding</scene> to <scene name='67/671725/Tp_i_in_the_presence_of_lps/4'>presence of LPS</scene>, making it <scene name='67/671725/Conformation_change/16'>more rigid and twisted</scene> than in the presence of water<ref name=Kushibiki>PMID:24389234</ref>. Moreover a docking model suggests the stability of the structure of TP-I is increased in the presence of LPS by the binding of the N and C termini of TP-I to LPS. The conformational change of TP-I seems to be crucial for its antimicrobial activity, since rearrangement of TP-I structure makes it more amphiphilic to negatively charged membrane of bacteria and fungus<ref name=Laederach>PMID:12369825</ref>. | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
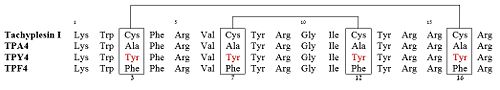
== Derivatives or Analogue == | == Derivatives or Analogue == | ||
Revision as of 08:21, 19 November 2020
| |||||||||||
References
- ↑ 1.0 1.1 1.2 1.3 1.4 Laederach A, Andreotti AH, Fulton DB. Solution and micelle-bound structures of tachyplesin I and its active aromatic linear derivatives. Biochemistry. 2002 Oct 15;41(41):12359-68. PMID:12369825
- ↑ 2.0 2.1 Chen, Yixin, et al. "RGD-Tachyplesin inhibits tumor growth." Cancer research 61.6 (2001): 2434-2438.
- ↑ 3.0 3.1 Saravanan R, Mohanram H, Joshi M, Domadia PN, Torres J, Ruedl C, Bhattacharjya S. Structure, activity and interactions of the cysteine deleted analog of tachyplesin-1 with lipopolysaccharide micelle: Mechanistic insights into outer-membrane permeabilization and endotoxin neutralization. Biochim Biophys Acta. 2012 Mar 23;1818(7):1613-1624. PMID:22464970 doi:10.1016/j.bbamem.2012.03.015
- ↑ Nakamura, Takanori, et al. "Tachyplesin, a class of antimicrobial peptide from the hemocytes of the horseshoe crab (Tachypleus tridentatus). Isolation and chemical structure." Journal of Biological Chemistry 263.32 (1988): 16709-16713
- ↑ 5.0 5.1 Kushibiki T, Kamiya M, Aizawa T, Kumaki Y, Kikukawa T, Mizuguchi M, Demura M, Kawabata SI, Kawano K. Interaction between tachyplesin I, an antimicrobial peptide derived from horseshoe crab, and lipopolysaccharide. Biochim Biophys Acta. 2014 Jan 2;1844(3):527-534. doi:, 10.1016/j.bbapap.2013.12.017. PMID:24389234 doi:http://dx.doi.org/10.1016/j.bbapap.2013.12.017
- ↑ 6.0 6.1 6.2 Hong, Jun, et al. "Mechanism of Tachyplesin I injury to bacterial membranes and intracellular enzymes, determined by laser confocal scanning microscopy and flow cytometry." Microbiological research (2014)
- ↑ Yonezawa A, Kuwahara J, Fujii N, Sugiura Y. Binding of tachyplesin I to DNA revealed by footprinting analysis: significant contribution of secondary structure to DNA binding and implication for biological action. Biochemistry. 1992 Mar 24;31(11):2998-3004. PMID:1372516
- ↑ Lipsky A, Cohen A, Ion A, Yedidia I. Genetic transformation of Ornithogalum via particle bombardment and generation of Pectobacterium carotovorum-resistant plants. Plant Sci. 2014 Nov;228:150-8. doi: 10.1016/j.plantsci.2014.02.002. Epub 2014 Feb, 12. PMID:25438795 doi:http://dx.doi.org/10.1016/j.plantsci.2014.02.002
- ↑ Lipsky A, Joshi JR, Carmi N, Yedidia I. Expression levels of antimicrobial peptide tachyplesin I in transgenic Ornithogalum lines affect the resistance to Pectobacterium infection. J Biotechnol. 2016 Nov 20;238:22-29. doi: 10.1016/j.jbiotec.2016.09.008. Epub, 2016 Sep 14. PMID:27639550 doi:http://dx.doi.org/10.1016/j.jbiotec.2016.09.008
- ↑ Ellerby, H. Michael, et al. "Anti-cancer activity of targeted pro-apoptotic peptides." Nature Medicine(1999)
Proteopedia Page Contributors and Editors (what is this?)
Shulamit Idzikowski, Janak Raj Joshi, Michal Harel, Alexander Berchansky, Joel L. Sussman, Angel Herraez, Jaime Prilusky