This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Megan Leaman/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 8: | Line 8: | ||
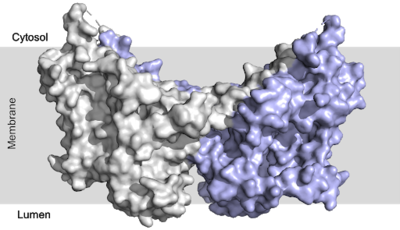
=== Conformation === | === Conformation === | ||
| - | There are three [https://en.wikipedia.org/wiki/Protein_domain domains] within the DGAT enzyme: cytosolic, transmembrane, and luminal. Most of the enzyme exists within the membrane with small portions peeking out into the cytosol of the cell or [https://en.wikipedia.org/wiki/Lumen_(anatomy) lumen] of the endoplasmic reticulum. DGAT exists as a [https://en.wikipedia.org/wiki/Protein_dimer homodimer] of two identical chains, A and B. The | + | There are three [https://en.wikipedia.org/wiki/Protein_domain domains] within the DGAT enzyme: cytosolic, transmembrane, and luminal. Most of the enzyme exists within the membrane with small portions peeking out into the cytosol of the cell or [https://en.wikipedia.org/wiki/Lumen_(anatomy) lumen] of the endoplasmic reticulum. DGAT exists as a [https://en.wikipedia.org/wiki/Protein_dimer homodimer] of two identical chains, A and B. The homodimer interface is stabilized in two different ways. First, the transmembrane region is stabilized through large <scene name='87/877557/Hydrophobic_interface/3'>hydrophobic interactions</scene> between the TM1 helices on each of the chains. The second is through extensive <scene name='87/877557/Hydrophilic_interface/5'>hydrogen bonding interactions</scene> between the two chains in the cytosolic domain. <ref name="Sui">PMID:32433611</ref> <ref name="Wang">PMID:32433610</ref> |
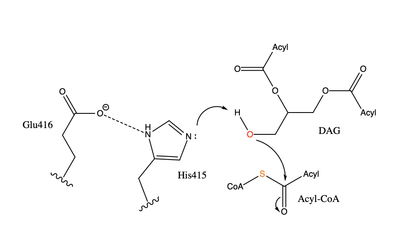
=== Active Site === | === Active Site === | ||
Revision as of 02:11, 21 April 2021
Human Diacylglycerol O-Transferase 1
| |||||||||||
References
- ↑ 1.0 1.1 Cases S, Smith SJ, Zheng YW, Myers HM, Lear SR, Sande E, Novak S, Collins C, Welch CB, Lusis AJ, Erickson SK, Farese RV Jr. Identification of a gene encoding an acyl CoA:diacylglycerol acyltransferase, a key enzyme in triacylglycerol synthesis. Proc Natl Acad Sci U S A. 1998 Oct 27;95(22):13018-23. PMID:9789033
- ↑ 2.0 2.1 2.2 2.3 2.4 2.5 2.6 2.7 2.8 Sui X, Wang K, Gluchowski NL, Elliott SD, Liao M, Walther TC, Farese RV Jr. Structure and catalytic mechanism of a human triacylglycerol-synthesis enzyme. Nature. 2020 May;581(7808):323-328. doi: 10.1038/s41586-020-2289-6. Epub 2020 May, 13. PMID:32433611 doi:http://dx.doi.org/10.1038/s41586-020-2289-6
- ↑ 3.0 3.1 Yen CL, Stone SJ, Koliwad S, Harris C, Farese RV Jr. Thematic review series: glycerolipids. DGAT enzymes and triacylglycerol biosynthesis. J Lipid Res. 2008 Nov;49(11):2283-301. doi: 10.1194/jlr.R800018-JLR200. Epub 2008, Aug 29. PMID:18757836 doi:http://dx.doi.org/10.1194/jlr.R800018-JLR200
- ↑ 4.0 4.1 4.2 4.3 Wang L, Qian H, Nian Y, Han Y, Ren Z, Zhang H, Hu L, Prasad BVV, Laganowsky A, Yan N, Zhou M. Structure and mechanism of human diacylglycerol O-acyltransferase 1. Nature. 2020 May;581(7808):329-332. doi: 10.1038/s41586-020-2280-2. Epub 2020 May, 13. PMID:32433610 doi:http://dx.doi.org/10.1038/s41586-020-2280-2
- ↑ Ransey E, Paredes E, Dey SK, Das SR, Heroux A, Macbeth MR. Crystal structure of the Entamoeba histolytica RNA lariat debranching enzyme EhDbr1 reveals a catalytic Zn(2+) /Mn(2+) heterobinucleation. FEBS Lett. 2017 Jul;591(13):2003-2010. doi: 10.1002/1873-3468.12677. Epub 2017, Jun 14. PMID:28504306 doi:http://dx.doi.org/10.1002/1873-3468.12677
Student Contributors
- Megan Leaman
- Grace Hall
- Karina Latsko