Introduction
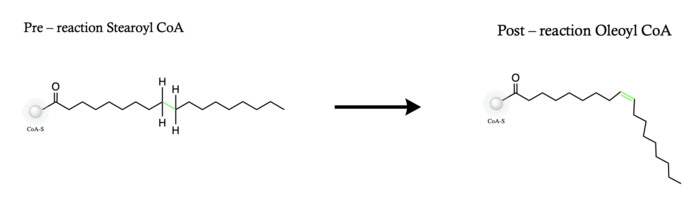
Stearyol CoA Desaturase (SCD1) functions as a lipogenic enzyme which is essential for fatty acid metabolism. SCD1 desaturates the sigma bond, within the 18-carbon acyl-CoA ligand, that attaches carbons 9 and 10[1]. The primary role of SCD1 is to catalyze the biosynthesis of monounsaturated fatty acids (MUFAs) via saturated acyl-CoAs with an acyl chain length of 14-19 carbons[2][3]. Variations of the monounsaturated fatty acids function as precursors for the biosynthesis of phospholipids, cholesteryl esters, and triglycerides; therefore, SCD1 is a promising candidate for drug targeting[1].
Biological Relevance
Absence or a deficit of SCD1 in the body is associated with obesity and insulin resistant which is a main cause of Type II diabetes type 2 diabetes[3]. Cancer sites in the body tend to show a much higher expression rate of SCD1[3]. Focusing on SCD1 as a drug target could lead to advancements in treatment of obesity, diabetes, and other metabolic diseases[1].
General Structure
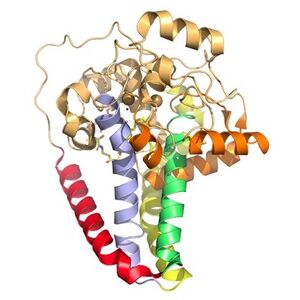
SCD1 is a protein transmembrane protein (4 helices in membrane, 8 helices in cytoplasm) that acquires electrons via an electron transport chain which includes cytochrome b5 reductase and cytochrome b5 . The electrons are transferred via a ternary complex and accepted by SCD1 by the iron metal ions[3]. SCD1 has 8 helices that are hydrophobic, 4 helices that are hydrophilic, and 3 helices that are amphipathic[1][3]. There are two Fe+2 metal ions within the structure of SCD1 that were determined by x-ray fluorescence chromatography x-ray fluorescense[3].
Necessary Structural Elements
Ligand Binding Pocket
The ligand binding pocket is a narrow tunnel that extends approximately 24 Å into the mostly hydrophobic interior of the protein. The ligand is stabilized by bending into a kinked conformation which creates a tight fit in the binding pocket tunnel, and by a hydrogen bond that occurs between the W258 side chain and the acyl carbonyl[1]. The kink in the tunnel is formed by the conserved residues, [1].7 which are stabilized by the hydrogen bond shared with Q143[1]. There are that interact with the substrate; the Fe2+ ions are coordinated by 9 histidine residues. One metal ion is coordinated by 4 histidines residues and a water molecule, and the other metal ion is coordinated by 5 histidine residues[1]. the ligand is seen to be in a eclipsed position, indicating it is in its post-reaction form. The histidines residues position the metal ions 6.4 Å apart[1].
Histidine Coordination
There are that together coordinate the metal ions[1]. One of the metal ions is coordinated by 4 histidines residues and a water molecule, and the other metal ion is coordinated by 5 histidine residues[1]. The histidines residues position the metal ions 6.4 Å apart[1].
Desaturation SIte
The ligand is desaturated at carbons 9 and 10[3]. The desaturation site of the ligand takes place inside the active site tunnel which enforces correct positioning of the substrate[1]. Before the reaction occurs, the ligand is in a gauche conformation at the desaturation site. This was determined by accidental usage of which allowed for binding of the substrate but prevented the reaction[3]. The product is in a cis conformation post-reaction. The product structure was determined using Fe2+ metal ions which allowed for the full reaction to take place[3]. The difference between the is the creation of a double bond, and the positioning of carbon 9 and 10 into a eclipsed position
Active Site Cap
The two conserved residues of the active site cap are . These two residues form a hydrogen bond creating a ridged barrier at the end of the active site to keep the ligand from moving during the reaction[1]. The active site cap is also used in determining the substrate length when entering the active site[1].
Catalytic Molecule
The of the SCD1 enzyme is a water molecule, coordinated by via hydrogen bonding[1]. The water molecule is 2.2 Å away from the Fe2+ metal ion molecule[1]. It interacts with the Fe2+ ion to make highly reactive radicals that are able to desaturate the highly stable carbon chain[4]. It is through the coordination of these ions by the histidines that the are able to be positioned within the vicinity of carbons 9 and 10 of the ligand[4].
Substrate Entering and Leaving
The substrate enters the active site through the active site tunnel and undergoes a to conform to the kinked shape of the tunnel[1]. Upon the substrate being converted to its final form, the product is laterally released from the protein due to a lateral egress of H1 and H2 caused by the deformation in the hydrogen bonding of the residues N 143 and T 257[1].
Proposed Mechanism
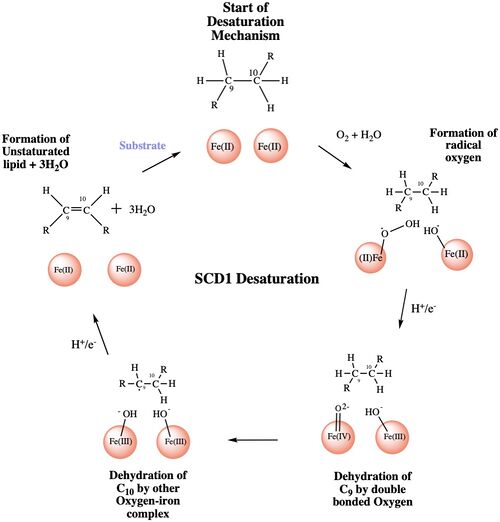
The mechanism used by the SCD1 enzyme is different than most desaturase enzymes because it does not use an oxo-bridge[3]. This mechanism begins with the addition of O2 and H2O which react with the Fe2+ ions to create oxygen radicals on the iron ions[3]. Then an electron/proton pair is brought in via the electron transport chain; this increases the oxidation of both iron ions, gets rid of the radicals, and creates an active iron-oxyl molecule[3]. The iron-oxyl molecule is reactive enough, due to the change in the oxidation state, to pull off the first hydrogen on carbon 9[4]. An unstable radical intermediate of the 18-carbon acyl-CoA ligand is formed which reacts with the other iron-O molecule, in the +3 state, to pull of the second hydrogen and form the final product (figure)[4]. Another electron/proton pair is brought in to create three H2O molecules and to take the iron ions back down to their original oxidation state of +2[4].
Regulation of Enzyme
The main function of SCD1 is to create the desaturated ligand that is used in the synthesis of cholesterol esters and triglycerides[3]. Based on this main function it has been confirmed that insulin and carbohydrate metabolism play a major role in the regulation of SCD1[5]. Along with insulin many other hormones have shown positive regulation of the SCD1 enzyme including T3, estradiol, and dexamethasone[5]. The carbohydrate metabolism can be altered by eating a fat-free, high carbohydrate diet with an increased intake of polyunsaturated fatty acids[5]. The will negatively regulate the mRNA expression of the SCD1 enzyme in the liver because high amounts of carbohydrates and polyunsaturated fatty acids decrease/inhibit the activity of the SCD1 enzyme[5].
References
- ↑ 1.00 1.01 1.02 1.03 1.04 1.05 1.06 1.07 1.08 1.09 1.10 1.11 1.12 1.13 1.14 1.15 1.16 1.17 1.18 Bai Y, McCoy JG, Levin EJ, Sobrado P, Rajashankar KR, Fox BG, Zhou M. X-ray structure of a mammalian stearoyl-CoA desaturase. Nature. 2015 Jun 22. doi: 10.1038/nature14549. PMID:26098370 doi:http://dx.doi.org/10.1038/nature14549
- ↑ Paton CM, Ntambi JM. Biochemical and physiological function of stearoyl-CoA desaturase. Am J Physiol Endocrinol Metab. 2009 Jul;297(1):E28-37. doi:, 10.1152/ajpendo.90897.2008. Epub 2008 Dec 9. PMID:19066317 doi:http://dx.doi.org/10.1152/ajpendo.90897.2008
- ↑ 3.00 3.01 3.02 3.03 3.04 3.05 3.06 3.07 3.08 3.09 3.10 3.11 3.12 Shen J, Wu G, Tsai AL, Zhou M. Structure and Mechanism of a Unique Diiron Center in Mammalian Stearoyl-CoA Desaturase. J Mol Biol. 2020 May 27. pii: S0022-2836(20)30367-3. doi:, 10.1016/j.jmb.2020.05.017. PMID:32470559 doi:http://dx.doi.org/10.1016/j.jmb.2020.05.017
- ↑ 4.0 4.1 4.2 4.3 4.4 doi: https://dx.doi.org/10.1021/acscatal.9b00456
- ↑ 5.0 5.1 5.2 5.3 Ntambi JM. The regulation of stearoyl-CoA desaturase (SCD). Prog Lipid Res. 1995;34(2):139-50. doi: 10.1016/0163-7827(94)00010-j. PMID:7480063 doi:http://dx.doi.org/10.1016/0163-7827(94)00010-j