We apologize for Proteopedia being slow to respond. For the past two years, a new implementation of Proteopedia has been being built. Soon, it will replace this 18-year old system. All existing content will be moved to the new system at a date that will be announced here.
User:Kaitlyn Roberts/Sandbox 2
From Proteopedia
(Difference between revisions)
| Line 9: | Line 9: | ||
== Structure == | == Structure == | ||
=== Tertiary Structure === | === Tertiary Structure === | ||
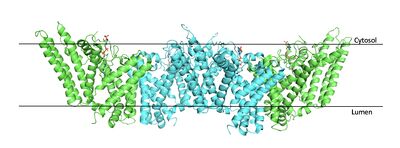
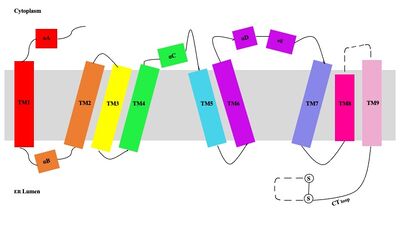
| - | [[Image:Tetramerlabels.jpeg| | + | [[Image:Tetramerlabels.jpeg|400 px|right|thumb|'''Figure 2.''' Tetramer unit of SOAT shown with membrane markers]] The overall structure of the enzyme is a <scene name='87/877559/Tetramer/10'>tetramer</scene> structure or a dimer of dimers. The functional building block of SOAT is a <scene name='87/877559/Dimer/3'>dimer</scene> which is made up of two identical <scene name='87/877559/Monomer/5'>monomer</scene> units. The residues that form the dimer interface are mostly hydrophobic and interact with each other in a shape-complementary manner. Mutating residues within the dimer interface reduced the dimers to monomer fractions, indicating that the dimeric architecture is important for the activity of the enzyme. Each monomer is organized into 9 <scene name='87/877559/Helices_1-9/4'>transmembrane helices</scene>. [[Image:Helicesdiagram1.jpeg|400 px|right|thumb|'''Figure 3.''' Labeled helices of SOAT within the membrane]]The dimerization of SOAT is mainly mediated by extensive [https://en.wikipedia.org/wiki/Van_der_Waals_force van der Waals interactions] between TM1 in one protomer and the [https://en.wikipedia.org/wiki/Lumen_(anatomy) lumenal segment] of TM6 and the [https://en.wikipedia.org/wiki/Cytosol cytosolic segment] of TM9 in the other. TM1, TM5, TM6 and TM9 from the two protomers enclose a deep hydrophobic pocket that is open to the lumenal side. Numerous hydrophobic residues on TM6 and TM9 from one protomer contact those on TM1 from the other protomer. On the intracellular side, hydrophobic residues on IH1 of each protomer interact with each other to stabilize the dimer. <ref name="Qian">PMID:32433614</ref> |
| - | + | ||
=== Tunnel System === | === Tunnel System === | ||
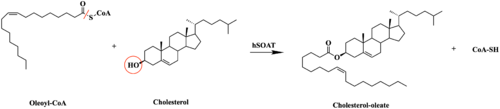
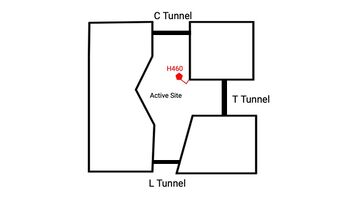
| - | [[Image:Tunnels2.jpg| | + | A main structural element of this enzyme is the tunnel systems. [[Image:Tunnels2.jpg|350 px|right|thumb|'''Figure 4.''' Tunnel system of SOAT]] There are 3 main tunnels in each monomer: the cytosolic (C) tunnel opening to the cytosol, the transmembrane(T) tunnel opening to the membrane, and the lumenal(L) tunnel opens to the lumen. The C tunnel opens to the cytosol of the cell and is the entrance site for the Acyl CoA into the active site. Surface representations of SOAT indicate that there are 2 alpha helices that block the entrance to the C tunnel, therefore a conformational change needs to occur to move the 2 helices so the substrate can enter the tunnel. The T tunnel opens into the membrane and is where cholesterol enters to have access to the active site. The two substrates are catalyzed by the H460 in the active site to form the cholesteryl ester. The products then leave via different pathways. The CoA-SH in the C tunnel leaves via that tunnel and is released back into the cytosol. The cholesteryl ester then leaves via either the T tunnel into the membrane or through the L tunnel into the lumen of the cell. <ref name="Qian" /> |
| - | + | ||
=== Active Site === | === Active Site === | ||
Revision as of 20:15, 24 April 2021
Human Sterol O-acyltransferase
| |||||||||||
References
- ↑ 1.0 1.1 1.2 Guan C, Niu Y, Chen SC, Kang Y, Wu JX, Nishi K, Chang CCY, Chang TY, Luo T, Chen L. Structural insights into the inhibition mechanism of human sterol O-acyltransferase 1 by a competitive inhibitor. Nat Commun. 2020 May 18;11(1):2478. doi: 10.1038/s41467-020-16288-4. PMID:32424158 doi:http://dx.doi.org/10.1038/s41467-020-16288-4
- ↑ 2.0 2.1 2.2 Qian H, Zhao X, Yan R, Yao X, Gao S, Sun X, Du X, Yang H, Wong CCL, Yan N. Structural basis for catalysis and substrate specificity of human ACAT1. Nature. 2020 May;581(7808):333-338. doi: 10.1038/s41586-020-2290-0. Epub 2020 May, 13. PMID:32433614 doi:http://dx.doi.org/10.1038/s41586-020-2290-0
- ↑ 3.0 3.1 Bhattacharyya R, Kovacs DM. ACAT inhibition and amyloid beta reduction. Biochim Biophys Acta. 2010 Aug;1801(8):960-5. doi: 10.1016/j.bbalip.2010.04.003. , Epub 2010 Apr 14. PMID:20398792 doi:http://dx.doi.org/10.1016/j.bbalip.2010.04.003
- ↑ 4.0 4.1 Huttunen HJ, Kovacs DM. ACAT as a drug target for Alzheimer's disease. Neurodegener Dis. 2008;5(3-4):212-4. doi: 10.1159/000113705. Epub 2008 Mar 6. PMID:18322393 doi:http://dx.doi.org/10.1159/000113705
- ↑ Chang C, Dong R, Miyazaki A, Sakashita N, Zhang Y, Liu J, Guo M, Li BL, Chang TY. Human acyl-CoA:cholesterol acyltransferase (ACAT) and its potential as a target for pharmaceutical intervention against atherosclerosis. Acta Biochim Biophys Sin (Shanghai). 2006 Mar;38(3):151-6. doi:, 10.1111/j.1745-7270.2006.00154.x. PMID:16518538 doi:http://dx.doi.org/10.1111/j.1745-7270.2006.00154.x
- ↑ Ayyagari VN, Wang X, Diaz-Sylvester PL, Groesch K, Brard L. Assessment of acyl-CoA cholesterol acyltransferase (ACAT-1) role in ovarian cancer progression-An in vitro study. PLoS One. 2020 Jan 24;15(1):e0228024. doi: 10.1371/journal.pone.0228024., eCollection 2020. PMID:31978092 doi:http://dx.doi.org/10.1371/journal.pone.0228024
Student Contributors
- Kylie Pfeifer
- Stephanie Pellegrino
- Kaitlyn Roberts