This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Betsy Johns/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 30: | Line 30: | ||
=== Oleoyl-CoA Binding === | === Oleoyl-CoA Binding === | ||
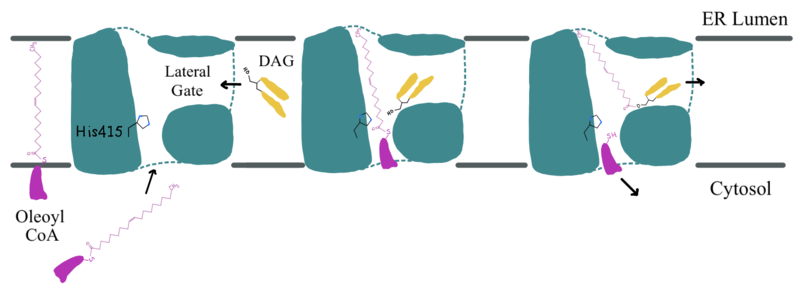
| - | Oleoyl-CoA enters DGAT1’s active site through a channel on the cytosolic side of the membrane. In order for the channel to widen and accommodate the fatty acid tail of the Oleoyl-CoA, <scene name='87/877512/Dgat_aligned2/16'>His415</scene> must flip from facing the cytosolic side to facing the lumenal side. It is speculated that the delta sulfur of the Met434 is involved in hydrogen bonding interactions with the catalytic His415 while it is flipped toward the cytosolic side of DGAT1. The His415 needs to break these speculated hydrogen bonds with Met434 to flip toward the lumenal side of DGAT1. Once <scene name='88/880292/Acyl_coa/2'>Oleoyl-CoA</scene> is bound in the active site, residues Asn378, Gln437, Met434, and Gln465 stabilize the fatty acid tail within the cytosolic channel of the active site, while residues His415 and Gln416 are directly involved within the catalytic mechanism of DGAT1. | + | Oleoyl-CoA enters DGAT1’s active site through a channel on the cytosolic side of the membrane. In order for the channel to widen and accommodate the fatty acid tail of the Oleoyl-CoA, <scene name='87/877512/Dgat_aligned2/16'>His415</scene> must flip from facing the cytosolic side to facing the lumenal side. It is speculated that the delta sulfur of the Met434 is involved in hydrogen bonding interactions with the catalytic His415 while it is flipped toward the cytosolic side of DGAT1 <ref name="Sui">PMID: 32433611</ref>. The His415 needs to break these speculated hydrogen bonds with Met434 to flip toward the lumenal side of DGAT1. Once <scene name='88/880292/Acyl_coa/2'>Oleoyl-CoA</scene> is bound in the active site, residues Asn378, Gln437, Met434, and Gln465 stabilize the fatty acid tail within the cytosolic channel of the active site, while residues His415 and Gln416 are directly involved within the catalytic mechanism of DGAT1 <ref name="Sui">PMID: 32433611</ref>. |
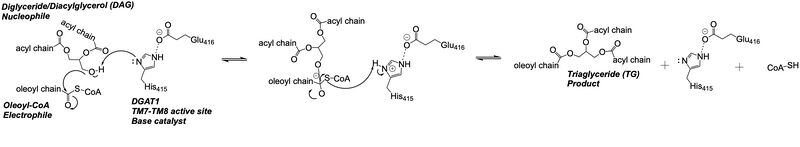
=== DAG Binding === | === DAG Binding === | ||
Revision as of 19:41, 26 April 2021
Diacylglycerol acyltransferase 1, DGAT1, synthesizes triacylglycerides
| |||||||||||
References
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 1.6 Wang L, Qian H, Nian Y, Han Y, Ren Z, Zhang H, Hu L, Prasad BVV, Laganowsky A, Yan N, Zhou M. Structure and mechanism of human diacylglycerol O-acyltransferase 1. Nature. 2020 May;581(7808):329-332. doi: 10.1038/s41586-020-2280-2. Epub 2020 May, 13. PMID:32433610 doi:http://dx.doi.org/10.1038/s41586-020-2280-2
- ↑ 2.0 2.1 2.2 2.3 2.4 2.5 2.6 2.7 2.8 Sui X, Wang K, Gluchowski NL, Elliott SD, Liao M, Walther TC, Farese RV Jr. Structure and catalytic mechanism of a human triacylglycerol-synthesis enzyme. Nature. 2020 May;581(7808):323-328. doi: 10.1038/s41586-020-2289-6. Epub 2020 May, 13. PMID:32433611 doi:http://dx.doi.org/10.1038/s41586-020-2289-6
- ↑ 3.0 3.1 3.2 Ma D, Wang Z, Merrikh CN, Lang KS, Lu P, Li X, Merrikh H, Rao Z, Xu W. Crystal structure of a membrane-bound O-acyltransferase. Nature. 2018 Oct;562(7726):286-290. doi: 10.1038/s41586-018-0568-2. Epub 2018 Oct, 3. PMID:30283133 doi:http://dx.doi.org/10.1038/s41586-018-0568-2
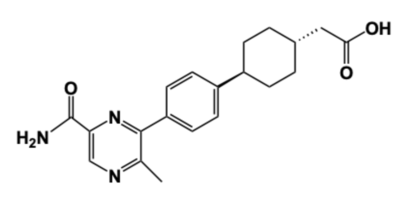
- ↑ 4.0 4.1 4.2 4.3 Denison H, Nilsson C, Lofgren L, Himmelmann A, Martensson G, Knutsson M, Al-Shurbaji A, Tornqvist H, Eriksson JW. Diacylglycerol acyltransferase 1 inhibition with AZD7687 alters lipid handling and hormone secretion in the gut with intolerable side effects: a randomized clinical trial. Diabetes Obes Metab. 2014 Apr;16(4):334-43. doi: 10.1111/dom.12221. Epub 2013 Oct, 31. PMID:24118885 doi:http://dx.doi.org/10.1111/dom.12221
- ↑ 5.0 5.1 Stephen J, Vilboux T, Haberman Y, Pri-Chen H, Pode-Shakked B, Mazaheri S, Marek-Yagel D, Barel O, Di Segni A, Eyal E, Hout-Siloni G, Lahad A, Shalem T, Rechavi G, Malicdan MC, Weiss B, Gahl WA, Anikster Y. Congenital protein losing enteropathy: an inborn error of lipid metabolism due to DGAT1 mutations. Eur J Hum Genet. 2016 Aug;24(9):1268-73. doi: 10.1038/ejhg.2016.5. Epub 2016 Feb , 17. PMID:26883093 doi:http://dx.doi.org/10.1038/ejhg.2016.5
- ↑ Ma D, Wang Z, Merrikh CN, Lang KS, Lu P, Li X, Merrikh H, Rao Z, Xu W. Crystal structure of a membrane-bound O-acyltransferase. Nature. 2018 Oct;562(7726):286-290. doi: 10.1038/s41586-018-0568-2. Epub 2018 Oct, 3. PMID:30283133 doi:http://dx.doi.org/10.1038/s41586-018-0568-2
Student Contributors
- Betsy Johns
- Elise Wang
- Tyler Bihasa