We apologize for Proteopedia being slow to respond. For the past two years, a new implementation of Proteopedia has been being built. Soon, it will replace this 18-year old system. All existing content will be moved to the new system at a date that will be announced here.
1lpa
From Proteopedia
(Difference between revisions)
| Line 3: | Line 3: | ||
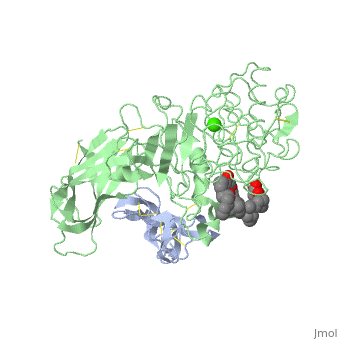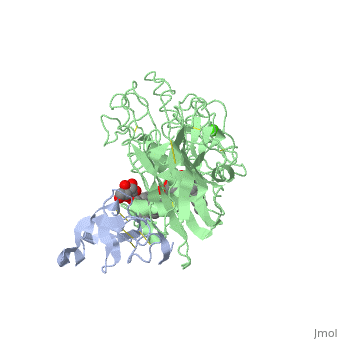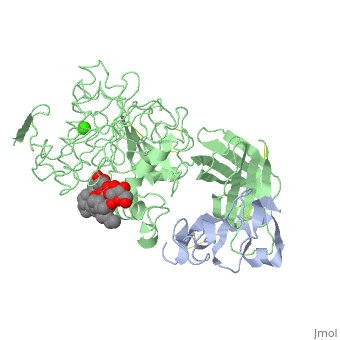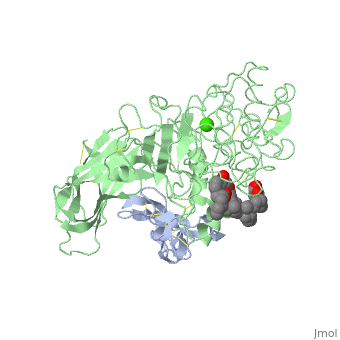
<StructureSection load='1lpa' size='340' side='right'caption='[[1lpa]], [[Resolution|resolution]] 3.04Å' scene=''> | <StructureSection load='1lpa' size='340' side='right'caption='[[1lpa]], [[Resolution|resolution]] 3.04Å' scene=''> | ||
== Structural highlights == | == Structural highlights == | ||
| - | <table><tr><td colspan='2'>[[1lpa]] is a 2 chain structure with sequence from [ | + | <table><tr><td colspan='2'>[[1lpa]] is a 2 chain structure with sequence from [https://en.wikipedia.org/wiki/Human Human] and [https://en.wikipedia.org/wiki/Pig Pig]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1LPA OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=1LPA FirstGlance]. <br> |
| - | </td></tr><tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat"><scene name='pdbligand=BNG:B-NONYLGLUCOSIDE'>BNG</scene>, <scene name='pdbligand=CA:CALCIUM+ION'>CA</scene>, <scene name='pdbligand=PLC:DIUNDECYL+PHOSPHATIDYL+CHOLINE'>PLC</scene></td></tr> | + | </td></tr><tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=BNG:B-NONYLGLUCOSIDE'>BNG</scene>, <scene name='pdbligand=CA:CALCIUM+ION'>CA</scene>, <scene name='pdbligand=PLC:DIUNDECYL+PHOSPHATIDYL+CHOLINE'>PLC</scene></td></tr> |
| - | <tr id='activity'><td class="sblockLbl"><b>Activity:</b></td><td class="sblockDat"><span class='plainlinks'>[ | + | <tr id='activity'><td class="sblockLbl"><b>Activity:</b></td><td class="sblockDat"><span class='plainlinks'>[https://en.wikipedia.org/wiki/Triacylglycerol_lipase Triacylglycerol lipase], with EC number [https://www.brenda-enzymes.info/php/result_flat.php4?ecno=3.1.1.3 3.1.1.3] </span></td></tr> |
| - | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[ | + | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=1lpa FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1lpa OCA], [https://pdbe.org/1lpa PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=1lpa RCSB], [https://www.ebi.ac.uk/pdbsum/1lpa PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=1lpa ProSAT]</span></td></tr> |
</table> | </table> | ||
== Function == | == Function == | ||
| - | [[ | + | [[https://www.uniprot.org/uniprot/COL_PIG COL_PIG]] Colipase is a cofactor of pancreatic lipase. It allows the lipase to anchor itself to the lipid-water interface. Without colipase the enzyme is washed off by bile salts, which have an inhibitory effect on the lipase. Enterostatin has a biological activity as a satiety signal. |
== Evolutionary Conservation == | == Evolutionary Conservation == | ||
[[Image:Consurf_key_small.gif|200px|right]] | [[Image:Consurf_key_small.gif|200px|right]] | ||
| Line 32: | Line 32: | ||
==See Also== | ==See Also== | ||
*[[Lipase 3D Structures|Lipase 3D Structures]] | *[[Lipase 3D Structures|Lipase 3D Structures]] | ||
| - | *[[Molecular Playground/Pancreatic Lipase|Molecular Playground/Pancreatic Lipase]] | ||
== References == | == References == | ||
<references/> | <references/> | ||
Revision as of 06:47, 18 August 2021
INTERFACIAL ACTIVATION OF THE LIPASE-PROCOLIPASE COMPLEX BY MIXED MICELLES REVEALED BY X-RAY CRYSTALLOGRAPHY
| |||||||||||