This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1tyf
From Proteopedia
(Difference between revisions)
| Line 3: | Line 3: | ||
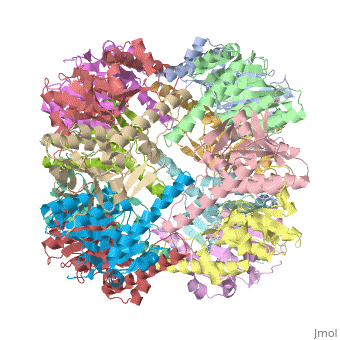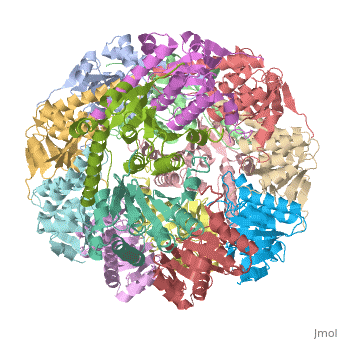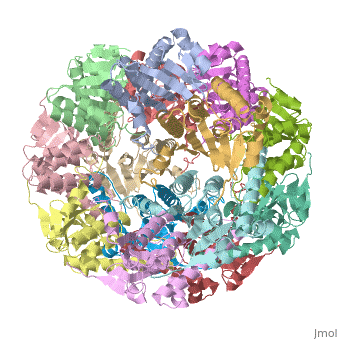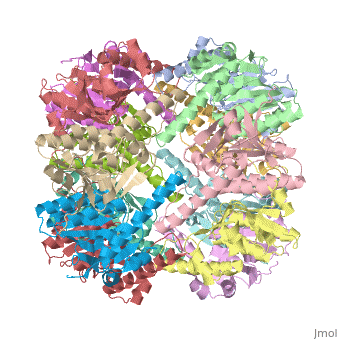
<StructureSection load='1tyf' size='340' side='right'caption='[[1tyf]], [[Resolution|resolution]] 2.30Å' scene=''> | <StructureSection load='1tyf' size='340' side='right'caption='[[1tyf]], [[Resolution|resolution]] 2.30Å' scene=''> | ||
== Structural highlights == | == Structural highlights == | ||
| - | <table><tr><td colspan='2'>[[1tyf]] is a 14 chain structure with sequence from [ | + | <table><tr><td colspan='2'>[[1tyf]] is a 14 chain structure with sequence from [https://en.wikipedia.org/wiki/"bacillus_coli"_migula_1895 "bacillus coli" migula 1895]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1TYF OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=1TYF FirstGlance]. <br> |
| - | </td></tr><tr id='activity'><td class="sblockLbl"><b>Activity:</b></td><td class="sblockDat"><span class='plainlinks'>[ | + | </td></tr><tr id='activity'><td class="sblockLbl"><b>Activity:</b></td><td class="sblockDat"><span class='plainlinks'>[https://en.wikipedia.org/wiki/Endopeptidase_Clp Endopeptidase Clp], with EC number [https://www.brenda-enzymes.info/php/result_flat.php4?ecno=3.4.21.92 3.4.21.92] </span></td></tr> |
| - | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[ | + | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=1tyf FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1tyf OCA], [https://pdbe.org/1tyf PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=1tyf RCSB], [https://www.ebi.ac.uk/pdbsum/1tyf PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=1tyf ProSAT]</span></td></tr> |
</table> | </table> | ||
== Function == | == Function == | ||
| - | [[ | + | [[https://www.uniprot.org/uniprot/CLPP_ECOLI CLPP_ECOLI]] Cleaves peptides in various proteins in a process that requires ATP hydrolysis. Has a chymotrypsin-like activity. Plays a major role in the degradation of misfolded proteins. May play the role of a master protease which is attracted to different substrates by different specificity factors such as ClpA or ClpX. |
== Evolutionary Conservation == | == Evolutionary Conservation == | ||
[[Image:Consurf_key_small.gif|200px|right]] | [[Image:Consurf_key_small.gif|200px|right]] | ||
Revision as of 09:34, 29 September 2021
THE STRUCTURE OF CLPP AT 2.3 ANGSTROM RESOLUTION SUGGESTS A MODEL FOR ATP-DEPENDENT PROTEOLYSIS
| |||||||||||