We apologize for Proteopedia being slow to respond. For the past two years, a new implementation of Proteopedia has been being built. Soon, it will replace this 18-year old system. All existing content will be moved to the new system at a date that will be announced here.
Sandbox Reserved 1706
From Proteopedia
(Difference between revisions)
| Line 8: | Line 8: | ||
==Introduction== | ==Introduction== | ||
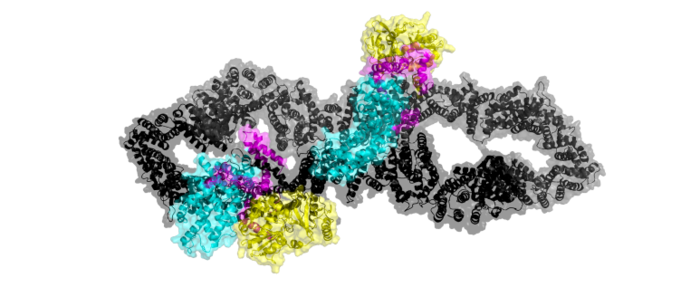
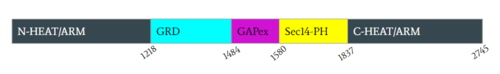
| - | [https://en.wikipedia.org/wiki/Neurofibromatosis_type_I Neurofibromatosis Type 1] is a genetic disorder caused by mutations in the tumor suppressor gene NF1 that codes for the [https://en.wikipedia.org/wiki/GTPase-activating_protein GTPase-activating protein] neurofibromin. Neurofibromin is closely involved in signaling pathways such as [https://en.wikipedia.org/wiki/MAPK/ERK_pathway MAPK/ERK], [https://en.wikipedia.org/wiki/PI3K/AKT/mTOR_pathway P13K/AKT/mTOR], and other cell signaling pathways that use [https://en.wikipedia.org/wiki/Ras_GTPase Ras] <ref name="Bergoug"> DOI:10.3390/cells9112365</ref>. Decreased activity of neurofibromin due to mutation can lead to tumor growth along nerves. As it is ubiquitous expression, NF1 misregulation can cause systemic tumor growth. Neurofibromin is localized to the [https://en.wikipedia.org/wiki/Cytosol cytosol] but is recruited to the [https://en.wikipedia.org/wiki/Cell_membrane plasma membrane] to inactivate [https://en.wikipedia.org/wiki/Ras_GTPase Ras]. The structure of neurofibromin was determined by high-resolution single | + | [https://en.wikipedia.org/wiki/Neurofibromatosis_type_I Neurofibromatosis Type 1] is a genetic disorder caused by mutations in the tumor suppressor gene NF1 that codes for the [https://en.wikipedia.org/wiki/GTPase-activating_protein GTPase-activating protein] neurofibromin. Neurofibromin is closely involved in signaling pathways such as [https://en.wikipedia.org/wiki/MAPK/ERK_pathway MAPK/ERK], [https://en.wikipedia.org/wiki/PI3K/AKT/mTOR_pathway P13K/AKT/mTOR], and other cell signaling pathways that use [https://en.wikipedia.org/wiki/Ras_GTPase Ras] <ref name="Bergoug"> DOI:10.3390/cells9112365</ref>. Decreased activity of neurofibromin due to mutation can lead to tumor growth along nerves. As it is ubiquitous expression, NF1 misregulation can cause systemic tumor growth. Neurofibromin is localized to the [https://en.wikipedia.org/wiki/Cytosol cytosol] but is recruited to the [https://en.wikipedia.org/wiki/Cell_membrane plasma membrane] to inactivate [https://en.wikipedia.org/wiki/Ras_GTPase Ras]. The structure of neurofibromin was determined by high-resolution single particle [https://en.wikipedia.org/wiki/Cryogenic_electron_microscopy cryo-EM]. These structures illustrated the domain architecture and conformational changes in neurofibromin, controlling Ras binding and inactivation. <ref name="Bergoug"> DOI:10.3390/cells9112365</ref><ref name="Bourne"> DOI:10.1038/39470</ref><ref name="Lupton"> DOI:10.1038/s41594-021-00687-2</ref><ref name="Naschberger"> DOI:10.1038/s41586-021-04024-x</ref> |
==Function== | ==Function== | ||
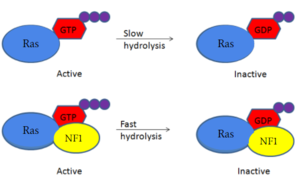
Neurofibromin is a [https://en.wikipedia.org/wiki/GTPase-activating_protein GTPase-activating protein] that binds to [https://en.wikipedia.org/wiki/Ras_GTPase Ras], a [https://en.wikipedia.org/wiki/GTPase GTPase], to increase its inherent hydrolysis of GTP to GDP (Figure 1). This inactivates the cell signaling of Ras until reactivated by [https://en.wikipedia.org/wiki/Guanosine_triphosphate GTP] exchange. Neurofibromin has two structural conformations, the open and closed conformation. Neurofibromin only binds to Ras in its open conformation. <ref name="Bourne"> DOI:10.1038/39470</ref><ref name="Lupton"> DOI:10.1038/s41594-021-00687-2</ref><ref name="Naschberger"> DOI:10.1038/s41586-021-04024-x</ref> [[Image:RasNeurofibrominMech.PNG|300px|right|thumb|Figure 1: Shifted rate of of Ras GTP hydrolysis when bound to neurofibromin. The speed of GTP hydrolysis is significantly increased when bound to neurofibromin. Ras is inactive when bound to GDP and active when bound to GTP ]] | Neurofibromin is a [https://en.wikipedia.org/wiki/GTPase-activating_protein GTPase-activating protein] that binds to [https://en.wikipedia.org/wiki/Ras_GTPase Ras], a [https://en.wikipedia.org/wiki/GTPase GTPase], to increase its inherent hydrolysis of GTP to GDP (Figure 1). This inactivates the cell signaling of Ras until reactivated by [https://en.wikipedia.org/wiki/Guanosine_triphosphate GTP] exchange. Neurofibromin has two structural conformations, the open and closed conformation. Neurofibromin only binds to Ras in its open conformation. <ref name="Bourne"> DOI:10.1038/39470</ref><ref name="Lupton"> DOI:10.1038/s41594-021-00687-2</ref><ref name="Naschberger"> DOI:10.1038/s41586-021-04024-x</ref> [[Image:RasNeurofibrominMech.PNG|300px|right|thumb|Figure 1: Shifted rate of of Ras GTP hydrolysis when bound to neurofibromin. The speed of GTP hydrolysis is significantly increased when bound to neurofibromin. Ras is inactive when bound to GDP and active when bound to GTP ]] | ||
Revision as of 23:54, 18 April 2022
| This Sandbox is Reserved from February 28 through September 1, 2022 for use in the course CH462 Biochemistry II taught by R. Jeremy Johnson at the Butler University, Indianapolis, USA. This reservation includes Sandbox Reserved 1700 through Sandbox Reserved 1729. |
To get started:
More help: Help:Editing |
Neurofibromin 1
| |||||||||||