This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Transcription-repair coupling factor
From Proteopedia
| Line 24: | Line 24: | ||
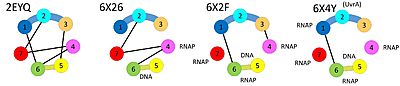
In the apo-structure, domains 3 and 4 are far apart from each other (connected by an extended linker), but they make direct contacts when Mfd is loaded on DNA. On the other hand, domains 4 and 5 make direct contacts in the apo-structure, but are far apart from each other (coonected by an extended linker) when Mfd is loaded on DNA. Finally, domain 2 and domain 7 bind to each other in the apo-state whereas domain 2 is available for binding UvrA in the DNA-bound state. | In the apo-structure, domains 3 and 4 are far apart from each other (connected by an extended linker), but they make direct contacts when Mfd is loaded on DNA. On the other hand, domains 4 and 5 make direct contacts in the apo-structure, but are far apart from each other (coonected by an extended linker) when Mfd is loaded on DNA. Finally, domain 2 and domain 7 bind to each other in the apo-state whereas domain 2 is available for binding UvrA in the DNA-bound state. | ||
| + | |||
== 3D Structures of Transcription-repair coupling factor == | == 3D Structures of Transcription-repair coupling factor == | ||
[[Transcription-repair coupling factor 3D structures]] | [[Transcription-repair coupling factor 3D structures]] | ||
| + | |||
</StructureSection> | </StructureSection> | ||
Revision as of 07:28, 1 May 2022
The bacterial transcription-repair coupling factor TRCF, also called Mfd translocase, is a DNA repair protein. It has a role in transcription-coupled repair, a subpathway of nucleotide excision repair (NER). Mfd recognizes stalled RNA polymerase (RNAP) and either restarts transcription or removes the stalled polymerase and recruits the NER proteins UvrA and UvrB.
Contents |
Function
Mfd has ATP hydrolysis activity, DNA binding sites and a UvrA binding sites. These three functions are inhibited in the isolated enzyme, but are activated when Mfd encounters stalled RNA polymerase (or through various mutations that remove inhibitory domains [1]). Mfd also contains an RNA interaction domain (RID) that binds to the beta subunit of RNAP.
Relationship to other enzymes
The N-terminal part of Mfd shows sequence similarity to UvrB, including in the domain of UvrB that interacts with UvrA. However, the conserved helicase motifs present in UvrB (responsible for binding and hydrolyzing ATP) are absent in that part of Mfd. Furthermore, the sequence segment known to fold as a beta hairpin in UvrB (involved in clamping down a single strand of DNA) seems absent. The C-terminal part of Mfd shows sequence similarity to SF1/SF2 helicases (UvrB is an example), containing conserved helicase/translocase motifs.
Structure and conformational change
| |||||||||||
References
- ↑ Selby CP. Mfd Protein and Transcription-Repair Coupling in Escherichia coli. Photochem Photobiol. 2017 Jan;93(1):280-295. doi: 10.1111/php.12675. Epub 2017, Jan 18. PMID:27864884 doi:http://dx.doi.org/10.1111/php.12675
- ↑ Deaconescu AM, Chambers AL, Smith AJ, Nickels BE, Hochschild A, Savery NJ, Darst SA. Structural basis for bacterial transcription-coupled DNA repair. Cell. 2006 Feb 10;124(3):507-20. PMID:16469698 doi:10.1016/j.cell.2005.11.045
- ↑ Brugger C, Zhang C, Suhanovsky MM, Kim DD, Sinclair AN, Lyumkis D, Deaconescu AM. Molecular determinants for dsDNA translocation by the transcription-repair coupling and evolvability factor Mfd. Nat Commun. 2020 Jul 27;11(1):3740. doi: 10.1038/s41467-020-17457-1. PMID:32719356 doi:http://dx.doi.org/10.1038/s41467-020-17457-1
- ↑ Shi J, Wen A, Zhao M, Jin S, You L, Shi Y, Dong S, Hua X, Zhang Y, Feng Y. Structural basis of Mfd-dependent transcription termination. Nucleic Acids Res. 2020 Nov 18;48(20):11762-11772. doi: 10.1093/nar/gkaa904. PMID:33068413 doi:http://dx.doi.org/10.1093/nar/gkaa904
- ↑ Kang JY, Llewellyn E, Chen J, Olinares PDB, Brewer J, Chait BT, Campbell EA, Darst SA. Structural basis for transcription complex disruption by the Mfd translocase. Elife. 2021 Jan 22;10. pii: 62117. doi: 10.7554/eLife.62117. PMID:33480355 doi:http://dx.doi.org/10.7554/eLife.62117
- ↑ The Storymorph Jmol scripts were used to create the interpolation shown in the morph. Coordinates available on Proteopedia
Created with the participation of Wayne Decatur
Proteopedia Page Contributors and Editors (what is this?)
Karsten Theis, Michal Harel, Alexander Berchansky, Wayne Decatur