This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
FirstGlance/How To Measure A Virus Capsid
From Proteopedia
(Difference between revisions)
| Line 19: | Line 19: | ||
(EEEV) consists of three distinct protein sequences, and a total of 720 chains. It has nearly 2 million non-hydrogen atoms (so about 3.6 million atoms including hydrogen). The structure [[6mx4]] determined by electron microscopy (4.4 Å resolution) has 12 chains, with instructions for constructing the entire capsid using 60 copies of the asymmetric unit. | (EEEV) consists of three distinct protein sequences, and a total of 720 chains. It has nearly 2 million non-hydrogen atoms (so about 3.6 million atoms including hydrogen). The structure [[6mx4]] determined by electron microscopy (4.4 Å resolution) has 12 chains, with instructions for constructing the entire capsid using 60 copies of the asymmetric unit. | ||
| - | [[FirstGlance in Jmol]] automatically constructs the capsid, and simplifies it to a subset of alpha carbon atoms small enough (not more than 25,000) to be analyzed efficiently in the FirstGlance and JSmol, both of which run in the Javascript of the web browser. FirstGlance offers a number of useful color schemes, including distance from center, colors that distinguish sequence-identical groups of chains, and amino-to-carboxy rainbow. When the structure is small enough that all alpha carbons can be displayed (not more than 250,000 alpha carbons; EEEV has 242,340), it can also be colored by charge, hydrophobic vs. polar, or evolutionary conservation. | + | [[FirstGlance in Jmol]] automatically constructs the capsid, and simplifies it to a subset of alpha carbon atoms small enough (not more than 25,000) to be analyzed efficiently in the FirstGlance and [[JSmol]], both of which run in the Javascript of the web browser. FirstGlance offers a number of useful color schemes, including distance from center, colors that distinguish sequence-identical groups of chains, and amino-to-carboxy rainbow. When the structure is small enough that all alpha carbons can be displayed (not more than 250,000 alpha carbons; EEEV has 242,340), it can also be colored by charge, hydrophobic vs. polar, or evolutionary conservation. |
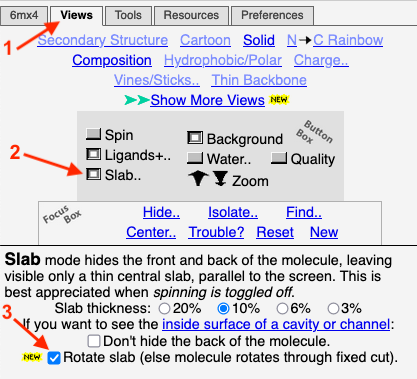
==Slab of the Capsid== | ==Slab of the Capsid== | ||
Revision as of 23:41, 26 July 2022
| |||||||||||