This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox reserved 1751
From Proteopedia
(Difference between revisions)
| Line 20: | Line 20: | ||
<div style="background-color:#fffaf0;"> | <div style="background-color:#fffaf0;"> | ||
== Publication Abstract from PubMed == | == Publication Abstract from PubMed == | ||
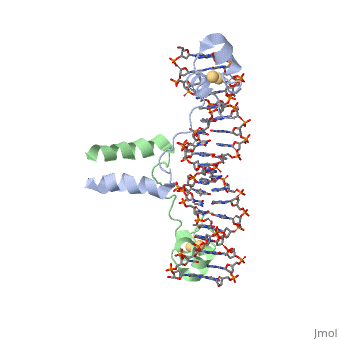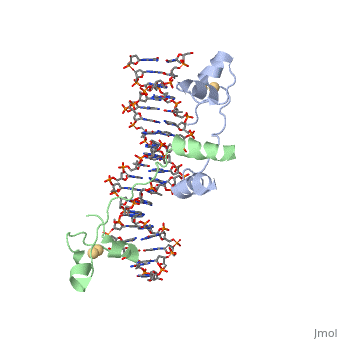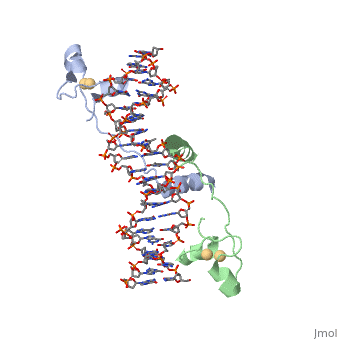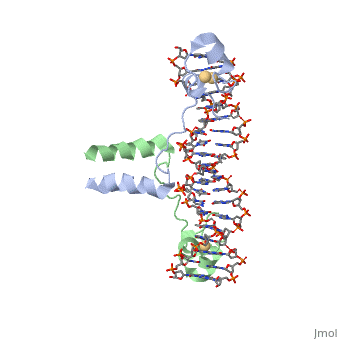
| - | A specific DNA complex of the 65-residue, N-terminal fragment of the yeast transcriptional activator, GAL4, has been analysed at 2.7 A resolution by X-ray crystallography. The protein binds as a <scene name='92/925551/Dimer_practice/1'>dimer</scene> to a symmetrical 17-base-pair sequence. Each subunit folds into three distinct modules: a compact, <scene name='92/925551/Dimer_practice/ | + | A specific DNA complex of the 65-residue, N-terminal fragment of the yeast transcriptional activator, GAL4, has been analysed at 2.7 A resolution by X-ray crystallography. The protein binds as a <scene name='92/925551/Dimer_practice/1'>dimer</scene> to a symmetrical 17-base-pair sequence. Each subunit folds into three distinct modules: a compact, <scene name='92/925551/Dimer_practice/12'>Metal Binding Domain</scene> (residues 8-40), an <scene name='92/925551/Dimer_practice/4'>Extended Linker</scene> (41-49), and an alpha-helical <scene name='92/925551/Dimer_practice/7'>Dimerization Element</scene> (50-64). A small, Zn(2+)-containing domain recognizes a conserved CCG triplet at each end of the site through direct contacts with the major groove. A short coiled-coil dimerization element imposes 2-fold symmetry. A segment of extended polypeptide chain links the metal-binding module to the dimerization element and specifies the length of the site. The relatively open structure of the complex would allow another protein to bind coordinately with GAL4. |
Gal4 <scene name='92/925551/Practice/1'>practice structure</scene> | Gal4 <scene name='92/925551/Practice/1'>practice structure</scene> | ||
Revision as of 02:55, 20 September 2022
DNA RECOGNITION BY GAL4: STRUCTURE OF A PROTEIN/DNA COMPLEX
| |||||||||||